General Information
-
DRAMP ID
- DRAMP20958
-
Peptide Name
- StigA16
-
Source
- Synthetic construct
-
Family
- Derived from scorpion venom peptide
-
Gene
- Not found
-
Sequence
- FFKLIPKLVKGLISAFK
-
Sequence Length
- 17
-
UniProt Entry
- No entry found
-
Protein Existence
- Synthetic
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial, Anti-Gram+, Anti-Gram-, Antifungal, Antiparasitic, Antiproliferative
-
Target Organism
-
- [Ref.29670004] Gram-positive bacteria: Staphylococcus aureus ATCC 29213 (MIC=2.34 μM), Staphylococcus epidermidis ATCC 122225 (MIC=9.38 μM), Enterococcus faecalis ATCC 4028 (MIC=1.17 μM);
- Gram-negative bacteria: Escherichia coli ATCC 25922 (MIC=2.34 μM), Enterobacter cloacae ATCC 13047 (MIC=9.38 μM), Pseudomonas aeruginosa ATCC 27853 (MIC=1.17 μM);
- yeasts: Candida albicans ATCC 90028 (MIC=4.69 μM), Candida krusei ATCC 6258 (MIC=9.38 μM), Candida glabrata ATCC 90030 (MIC=9.38 μM);
- StigA16 inhibit 100% of the epimastigote forms of Trypanosoma cruzi growth at 2.5 µM;
- HeLa cell line: IC50=13.01 µM
-
Hemolytic Activity
-
- [Ref.29670004] Hemolysis 0% at 1.17 [Ref.29670004] Hemolysis 0% at 1.17 μM, hemolysis 2% at 2.34 μM, hemolysis 10% at 4.69 μM, hemolysis 20% at 9.38 μM, hemolysis 35% at 18.75 μM, hemolysis 37% at 37.50 μM, hemolysis 40% at 75.00 μM against human red blood cells
-
Cytotoxicity
-
- [Ref.29670004] The cell viability of HeLa cell lines is 15%, 21%, 16%, 9%, 8%, 10% and 11% at peptide concentrations of 2, 4, 8, 10, 15, 20 and 40 μM.
- The cell viability of 3T3 cell lines is 69%, 77%, 62%, 47%, 21%, 15% and 15% at peptide concentratio
-
Binding Target
- Not found
Structure Information
-
Linear/Cyclic
- Linear
-
N-terminal Modification
- Free
-
C-terminal Modification
- Amidation
-
Nonterminal Modifications and Unusual Amino Acids
- None
-
Stereochemistry
- L
-
Structure
- Alpha helix
-
Structure Description
- The peptide adopts helical conformation with some random structure at the N- and C-terminals. In sodium phosphate buffer (PBS) and water they have a predominantly random structure, but in 20 mM sodium dodecyl sulfate (SDS) and 2,2,2-trifluoroethanol (TFE) they have a typical alpha-helix structure.
-
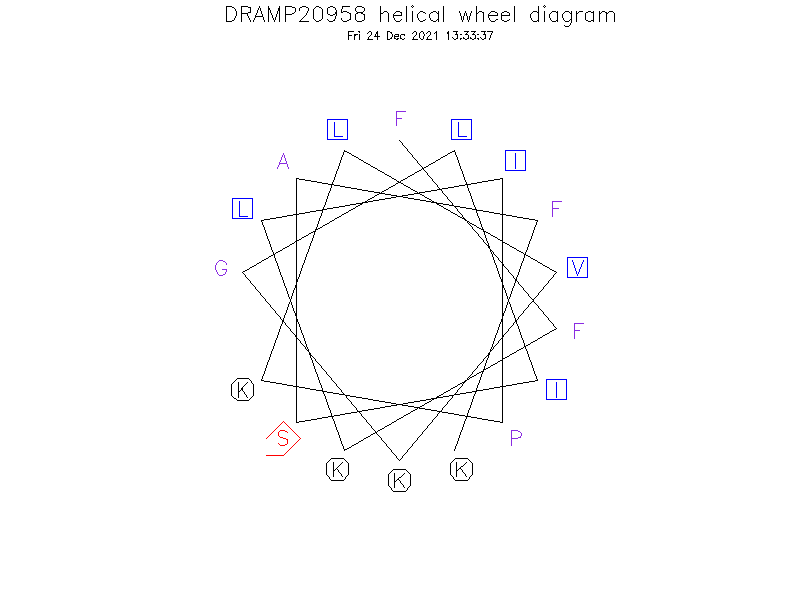
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP20958.
Physicochemical Information
-
Formula
- C99H161N21O19
Absent Amino Acids
- CDEHMNQRTWY
Common Amino Acids
- K
Mass
- 1949.5
PI
- 10.48
Basic Residues
- 4
Acidic Residues
- 0
Hydrophobic Residues
- 10
Net Charge
- +4
-
Boman Index
- 1473
Hydrophobicity
- 0.965
Aliphatic Index
- 137.65
Half Life
-
- Mammalian:1.1 hour
- Yeast:3 min
- E.coli:2 min
Extinction Coefficient Cystines
- 0
Absorbance 280nm
- 0
Polar Residues
- 2
DRAMP20958
Comments Information
Function
- Antibacterial activity against the Gram-positive and Gram-negative bacteria. Antifungal activity against yeasts.Antiparasitic Activity against epimastigote forms of Trypanosoma cruzi. Antiproliferative Activity against HeLa cells.
Literature Information
- ·Literature 1
-
Title
- Analogs of the Scorpion Venom Peptide Stigmurin: Structural Assessment, Toxicity, and Increased Antimicrobial Activity.
-
Pubmed ID
- 29670004
-
Reference
- Toxins (Basel). 2018 Apr 18;10(4). pii: E161.
-
Author
- Parente AMS, Daniele-Silva A, Furtado AA, Melo MA, Lacerda AF, Queiroz M, Moreno C, Santos E, Rocha HAO, Barbosa EG, Carvalho E, Silva-Júnior AA, Silva MS, Fernandes-Pedrosa MF.