General Information
-
DRAMP ID
- DRAMP21616
-
Peptide Name
- DRIM
-
Source
- Synthetic construct
-
Family
- Not found
-
Gene
- Not found
-
Sequence
- qqrkrkiwsilaplgttlvklvagig
-
Sequence Length
- 26
-
UniProt Entry
- No entry found
-
Protein Existence
- Synthetic form
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial, Anti-Gram+, Anti-Gram-
-
Target Organism
-
- [Ref.28921993] Gram-positive bacteria: Staphylococcus aureus (MIC = 32 μg/mL), Enterococcus faecalis (MIC = 32 μg/mL);
- Gram-negative bacteria: Escherichia coli (MIC = 32 μg/mL), Pseudomonas aeruginosa (MIC = 64 μg/mL)
-
Hemolytic Activity
-
- [Ref.28921993] 60.5%, 98.0%, 95.1%, 99.0%, 97.9%, 96.3%, 98.8% and 98.2% hemolysis against human red blood cells at 5, 7.5, 10, 15, 20, 25, 30 and 40 μg/ml.
-
Cytotoxicity
-
- [Ref.28921993] The toxicity of DRIM toward HEK293 and HeLa cells is much less than nonaarginine (R9) by use of flow cytometry and the peptide doesn't show any cytotoxicity at 2 μM.
-
Binding Target
- Not found
Structure Information
-
Linear/Cyclic
- Linear
-
N-terminal Modification
- Free
-
C-terminal Modification
- Amidation
-
Nonterminal Modifications and Unusual Amino Acids
- None
-
Stereochemistry
- D
-
Structure
- ①Disordered conformation in aqueous solutions [pure water (H₂O), phosphate buffer (PB, 10 mM), and phosphate buffer with high salt (NaF, 100 mM)]. ②50
-
Structure Description
- Our CD data show that all four linear peptides were generally disordered in aqueous solution, as expected, but prominent effects were seen regarding the impact of the staple.
-
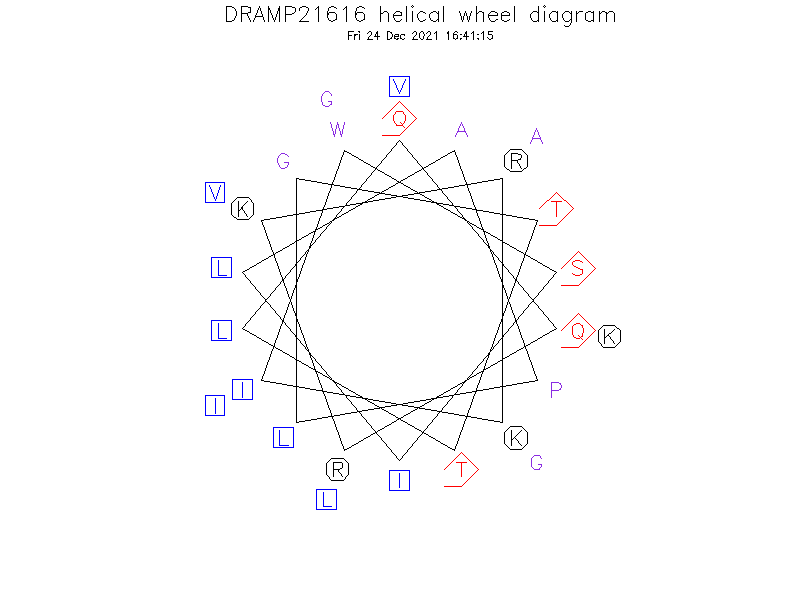
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP21616.
Physicochemical Information
-
Formula
- C₁₃₁H₂₂₈N₃₈O₃₂
Absent Amino Acids
- CDEFHMNY
Common Amino Acids
- L
Mass
- 2847.49
PI
- 12.02
Basic Residues
- 5
Acidic Residues
- 0
Hydrophobic Residues
- 12
Net Charge
- +5
-
Boman Index
- -1482
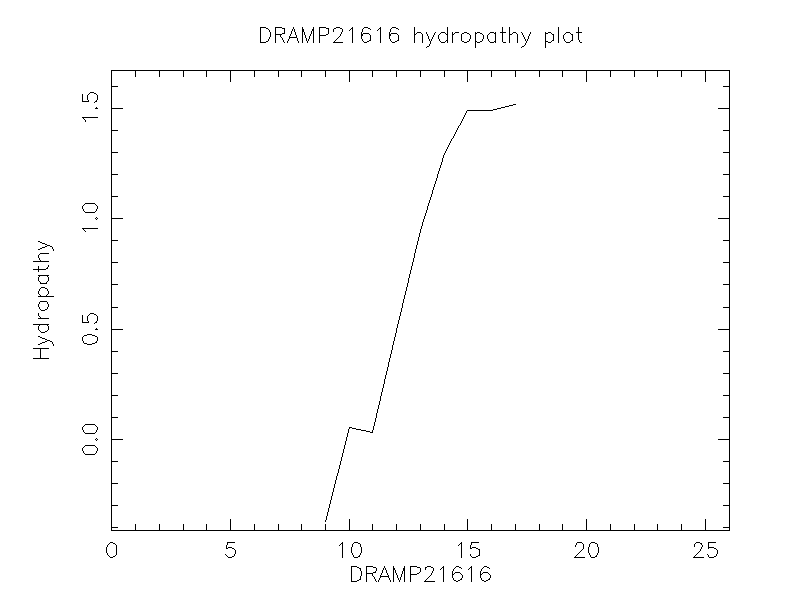
Hydrophobicity
- 0.273
Aliphatic Index
- 135
Half Life
-
- Mammalian:0.8 hour
- Yeast:10 min
- E.coli:10 hour
Extinction Coefficient Cystines
- 5500
Absorbance 280nm
- 220
Polar Residues
- 6
DRAMP21616
Comments Information
Function
- Antibacterial activity against Gram-positive and Gram-negative bacteria.
Literature Information
- ·Literature 1
-
Title
- Lactam-Stapled Cell-Penetrating Peptides: Cell Uptake and Membrane Binding Properties.
-
Pubmed ID
- 28921993
-
Reference
- J Med Chem. 2017 Oct 12;60(19):8071-8082. doi: 10.1021/acs.jmedchem.7b00813. Epub 2017 Sep 26.
-
Author
- Marco J Klein, Samuel Schmidt, Parvesh Wadhwani, Jochen Bürck, Johannes Reichert, Sergii Afonin, Marina Berditsch, Tim Schober, Roland Brock, Manfred Kansy, Anne S Ulrich