General Information
-
DRAMP ID
- DRAMP00107
-
Peptide Name
- Bacteriocin L-1077
-
Source
- Lactobacillus salivarius L-1077 (NRRL B-50053) (Gram-positive bacteria)
-
Family
- Belongs to the class IIa bacteriocin
-
Gene
- Not found
-
Sequence
- TNYGNGVGVPDAIMAGIIKLIFIFNIRQGYNFGKKAT
-
Sequence Length
- 37
-
UniProt Entry
- No entry found
-
Protein Existence
- Not found
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial, Anti-Gram+, Anti-Gram-
-
Target Organism
-
- Gram-negative bacteria:
Target Organism Activity Salmonella Enteritidis 1 MIC=0.19 ug/ml S. Enteritidis 4 MIC=0.38 ug/ml S. Enteritidis 204 MIC=0.38 ug/ml S. Enteritidis 237 MIC=0.19 ug/ml Salmonella Choleraesuis 434/4 MIC=0.38 ug/ml S. Choleraesuis 370 MIC=0.19 ug/ml Salmonella Typhimurium 483/6 MIC=0.19 ug/ml S. Typhimurium 44/8 MIC=0.19 ug/ml Salmonella Gallinarum-Pullorum MIC=0.19 ug/ml Escherichia coli HB 101 MIC=0.19 ug/ml E. coli-600 MIC=0.19 ug/ml E. coli O157:H7 Y-63 MIC=0.19 ug/ml E. coli O157:H7 G-35 MIC=0.19 ug/ml E. coli O157:H7 OD 30 MIC=0.19 ug/ml E. coli O157:H7 T lab 39 MIC=0.19 ug/ml Yersinia enterocolitica 03 MIC=0.76 ug/ml Y. enterocolitica 09 MIC=0.76 ug/ml Y. enterocolitica 11 MIC=0.76 ug/ml Citrobacter freundii MIC=0.76 ug/ml Klebsiella pneumoniae MIC=0.76 ug/ml Shigella dysenteriae MIC=0.76 ug/ml Yersinia pseudotuberculosis 4 MIC=0.76 ug/ml Y. pseudotuberculosis 99/14 MIC=0.76 µg/ml Pseudomonas aeruginosa 508 MIC=0.38 ug/ml Proteus mirabilis MIC=0.38 ug/ml Morganella morganii MIC=0.38 µg/ml Proteus vulgaris 7A MIC=1.5 ug/ml Campylobacter jejuni L-4 (MIC=0.09 ug/ml); - - Gram-positive bacteria:
Target Organism Activity Clostridium perfringens 13124 MIC=0.7 ug/ml Listeria monocytogenes A 9-72 MIC=0.19 ug/ml Staphylococcus aureus MIC=0.76 ug/ml Staphylococcus epidermidis MIC=0.76 ug/ml
- Gram-negative bacteria:
-
Hemolytic Activity
-
- No hemolysis information or data found in the reference(s) presented in this entry
-
Cytotoxicity
- No cytotoxicity information found in the reference(s) presented
-
Binding Target
- Not found
Structure Information
-
Linear/Cyclic
- Linear
-
N-terminal Modification
- Free
-
C-terminal Modification
- Free
-
Nonterminal Modifications and Unusual Amino Acids
- None
-
Stereochemistry
- L
-
Structure
- Not found
-
Structure Description
- Not found
-
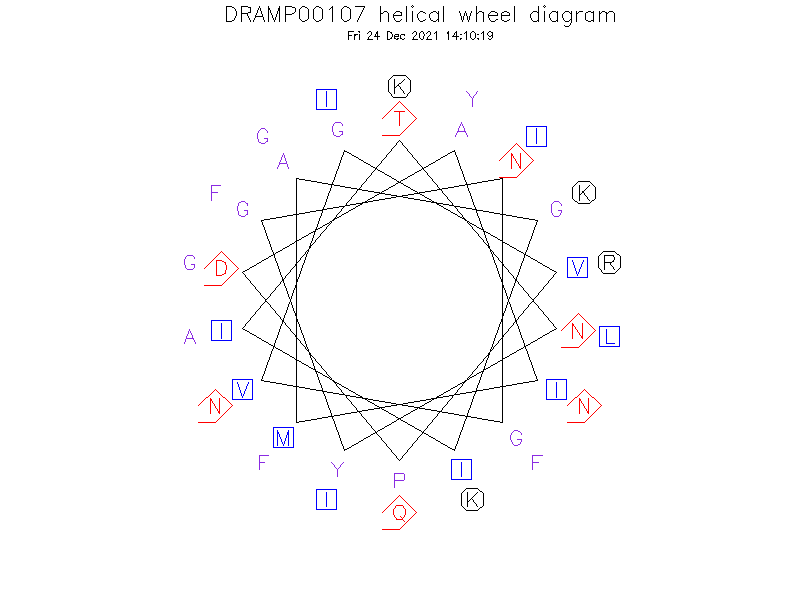
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP00107.
Physicochemical Information
-
Formula
- C185H290N48O49S
Absent Amino Acids
- CEHSW
Common Amino Acids
- GI
Mass
- 4002.69
PI
- 9.82
Basic Residues
- 4
Acidic Residues
- 1
Hydrophobic Residues
- 15
Net Charge
- +3
-
Boman Index
- -12.93
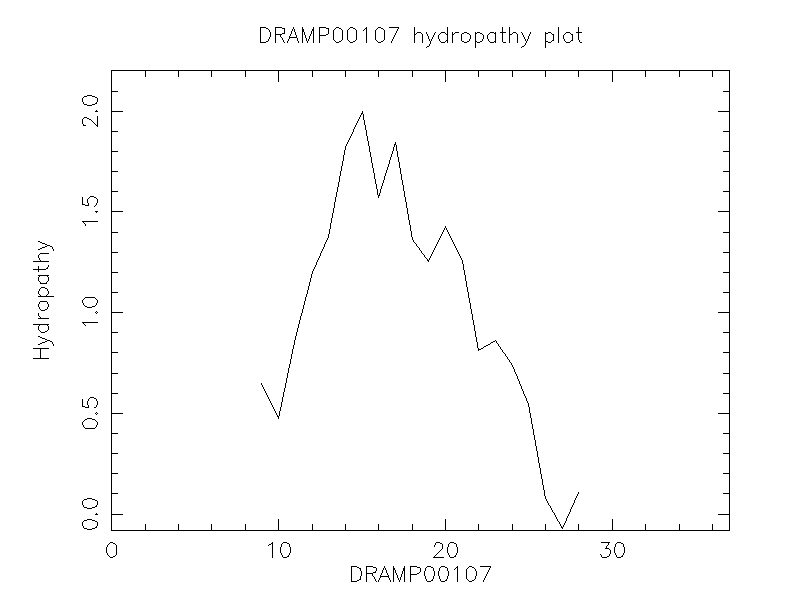
Hydrophobicity
- 0.262
Aliphatic Index
- 97.57
Half Life
-
- Mammalian:7.2 hour
- Yeast:>20 hour
- E.coli:>10 hour
Extinction Coefficient Cystines
- 2980
Absorbance 280nm
- 82.78
Polar Residues
- 14
DRAMP00107
Comments Information
Comment
- No comments found on DRAMP database
Literature Information
- ·Literature 1
-
Title
- Isolation of Lactobacillus salivarius 1077 (NRRL B-50053) and characterization of its bacteriocin, including the antimicrobial activity spectrum.
-
Pubmed ID
- 21378051
-
Reference
- Appl Environ Microbiol. 2011 Apr;77(8):2749-2754.
-
Author
- Svetoch EA, Eruslanov BV, Levchuk VP, Perelygin VV, Mitsevich EV, Mitsevich IP, Stepanshin J, Dyatlov I, Seal BS, Stern NJ.