General Information
-
DRAMP ID
- DRAMP00136
-
Peptide Name
- Enterocin E-760 (Bacteriocin)
-
Source
- Enterococcus sp. (Gram-positive bacteria)
-
Family
- Belongs to the class IIb bacteriocin
-
Gene
- Not found
-
Sequence
- NRWYCNSAAGGVGGAAVCGLAGYVGEAKENIAGEVRKGWGMAGGFTHNKACKSFPGSGWASG
-
Sequence Length
- 62
-
UniProt Entry
- P85147
-
Protein Existence
- Protein level
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial, Anti-Gram+, Anti-Gram-
-
Target Organism
-
- [Ref.18086839]Gram-positive bacteria:
Target Organism Activity Listeria monocytogenes 9-72 MIC=0.1 μg/ml Staphylococcus epidermidis MIC=1.6 μg/ml S. aureus (MIC=1.6 μg/ml); - - Gram-negative bacteria:
Target Organism Activity S. enterica serovar Enteritidis 1 MIC=0.2 μg/ml S. enterica serovar Enteritidis 4 MIC=0.4 μg/ml S. enterica serovar Enteritidis 204 MIC=0.2 μg/ml S. enterica serovar Enteritidis 237 MIC=0.2 μg/ml S. enterica serovar Choleraesuis 434/4 MIC=0.4 μg/ml S. enterica serovar Choleraesuis 370 MIC=0.4 μg/ml S. enterica serovar Typhimurium 383/60 MIC=0.4 μg/ml S. enterica serovar Typhimurium 320 MIC=0.2 μg/ml S. enterica serovar Gallinarum biovar Pullorum MIC=0.4 μg/ml Escherichia coli HB101 MIC=0.1 μg/ml E. coli C600 MIC=0.1 μg/ml E. coli O157:H7 Y-63 MIC=1.6 μg/ml E. coli O157:H7 G-3 MIC=1.6 μg/ml E. coli O157:H7 OD-3 MIC=1.6 μg/ml E. coli O157:H7 T-39 MIC=1.6 μg/ml Yersinia enterocolitica 03 MIC=0.1 μg/ml Y. enterocolitica 09 MIC=0.1 μg/ml Y. enterocolitica 04 MIC=0.1 μg/ml Y. pseudotuberculosis ser 4 MIC=3.2 μg/ml Y. pseudotuberculosis 14 MIC=3.2 μg/ml Citrobacter freundii MIC=1.6 μg/ml Klebsiella pneumoniae MIC=3.2 μg/ml Shigella dysenteriae MIC=0.1 μg/ml Campylobacter jejuni L4 MIC=0.1 μg/ml Campylobacter P 1-R MIC=0.2 μg/ml Campylobacter P-45 MIC=0.4 μg/ml Campylobacter lari R MIC=0.2 μg/ml Campylobacter LB 6-R MIC=0.1 μg/ml Campylobacter C/D MIC=0.2 μg/ml
- [Ref.18086839]Gram-positive bacteria:
-
Hemolytic Activity
-
- No hemolysis information or data found in the reference(s) presented in this entry
-
Cytotoxicity
- No cytotoxicity information found in the reference(s) presented
-
Binding Target
- Not found
Structure Information
-
Linear/Cyclic
- Linear
-
N-terminal Modification
- Free
-
C-terminal Modification
- Free
-
Nonterminal Modifications and Unusual Amino Acids
- None
-
Stereochemistry
- L
-
Structure
- Not found
-
Structure Description
- Not found
-
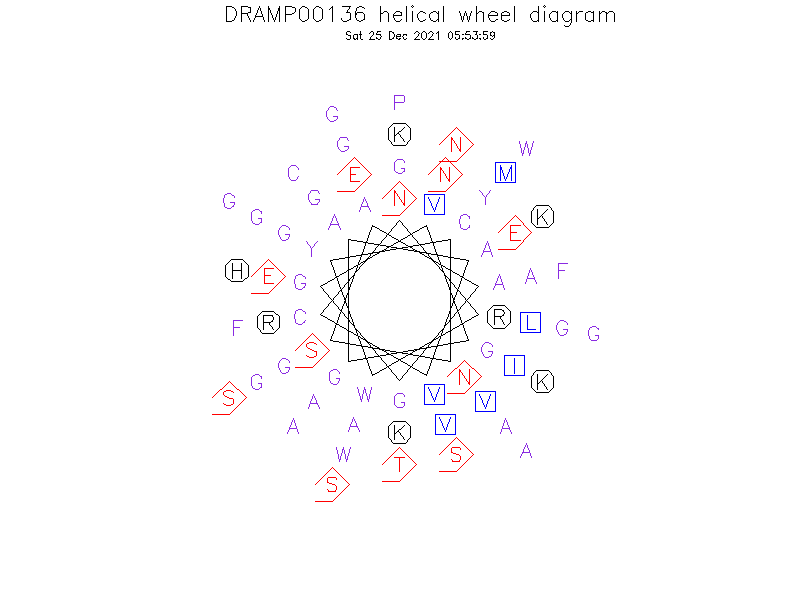
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP00136.
Physicochemical Information
-
Formula
- C269H403N81O80S4
Absent Amino Acids
- DQ
Common Amino Acids
- G
Mass
- 6179.89
PI
- 8.99
Basic Residues
- 7
Acidic Residues
- 3
Hydrophobic Residues
- 21
Net Charge
- +4
-
Boman Index
- -42.8
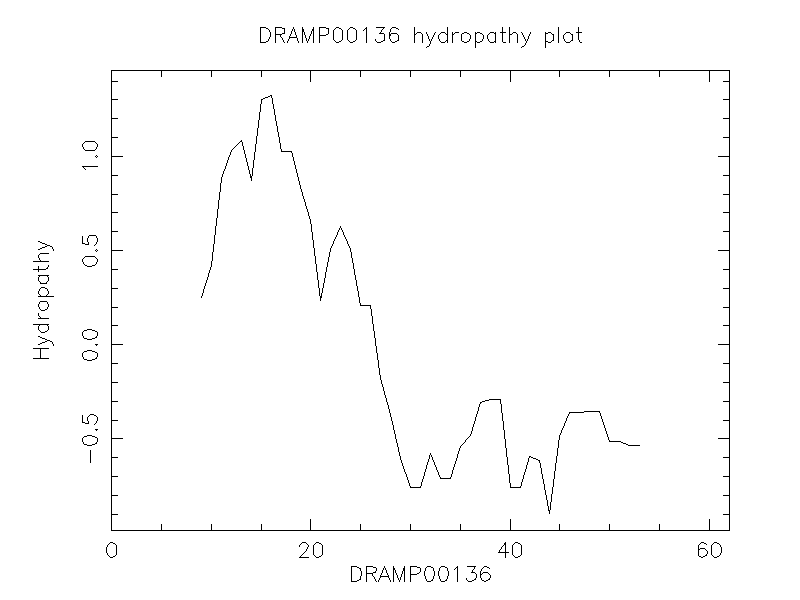
Hydrophobicity
- -0.177
Aliphatic Index
- 47.42
Half Life
-
- Mammalian:1.4 hour
- Yeast:3 min
- E.coli:>10 hour
Extinction Coefficient Cystines
- 19605
Absorbance 280nm
- 321.39
Polar Residues
- 29
DRAMP00136
Comments Information
Function
- The enterocin demonstrated thermostability by retaining activity after 5 min at 100 degrees C and was stable at pH values between 5.0 and 8.7. However, activity was lost below pH 3.0 and above pH 9.5. No hemolytic activity.
Literature Information
- ·Literature 1
-
Title
- Isolation and purification of enterocin E-760 with broad antimicrobial activity against gram-positive and Gram-negative bacteria.
-
Pubmed ID
- 18086839
-
Reference
- Antimicrob Agents Chemother. 2008 Mar;52(3):1094-1100.
-
Author
- Line JE, Svetoch EA, Eruslanov BV, Perelygin VV, Mitsevich EV, Mitsevich IP, Levchuk VP, Svetoch OE, Seal BS, Siragusa GR, Stern NJ.