General Information
-
DRAMP ID
- DRAMP00173
-
Peptide Name
- Leucocyclicin Q (Bacteriocin)
-
Source
- Leuconostoc mesenteroides TK41401 (Gram-positive bacteria)
-
Family
- Belongs to the class IIc bacteriocin
-
Gene
- lcyQ
-
Sequence
- LVNQLGISKSLANTILGAIAVGNLASWVLALVPGPGWATKAALATAETIVKHEGKAAAIAW
-
Sequence Length
- 61
-
UniProt Entry
- G5ELQ0
-
Protein Existence
- Predicted
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial, Anti-Gram+
-
Target Organism
-
- Gram-positive bacteria:
Target Organism Activity Lactococcus lactis ssp. lactis JCM 7638 MIC=0.038 µM L. lactis ssp. lactis ATCC 19435 MIC=0.038 µM L. sp. QU 12 MIC=1.203 µM Lactobacillus sakei ssp. sakei JCM 1157 MIC=0.038 µM Weissella cibaria JCM 12495 MIC=2.405 µM W. paramesenteroides JCM 9890 MIC=0.075 µM Pediococcus pentosaceus JCM 5885 MIC=2.405 µM P. dextrinicus JCM 5887 MIC=0.038 µM Enterococcus faecium JCM 5804 MIC=0.038 µM E. faecalis JCM 5803 MIC=0.15 µM Streptococcus salivarius JCM 5707 MIC=2.405 µM S. bovis JCM 5802 MIC=0.601 µM S. mutans JCM 5705 MIC=19.25 µM Bacillus coagulans JCM 2257 MIC=0.075 µM B. subtilis ssp. subtilis JCM 1465 MIC=0.601 µM B. cereus JCM 2152 MIC=2.405 µM Kocuria rhizophila NBRC 12708 MIC=9.623 µM Listeria innocua ATCC 33090 MIC=0.601 µM Leuconostoc mesenteroides ssp. mes. JCM 6124 MIC=0.3 µM Escherichia coli JM109 MIC=38.49 µM E. coli NBRC 3301 MIC=38.49 µM
- Gram-positive bacteria:
-
Hemolytic Activity
-
- No hemolysis information or data found in the reference(s) presented in this entry
-
Cytotoxicity
- No cytotoxicity information found in the reference(s) presented
-
Binding Target
- Not found
Structure Information
-
Linear/Cyclic
- Cyclic
-
N-terminal Modification
- Cyclization (N termini to C termini)
-
C-terminal Modification
- Cyclization (C termini to N termini)
-
Nonterminal Modifications and Unusual Amino Acids
- The whole peptide has a cyclic structure in which N and C termini are bound to each other.
-
Stereochemistry
- L
-
Structure
- Not found
-
Structure Description
- Not found
-
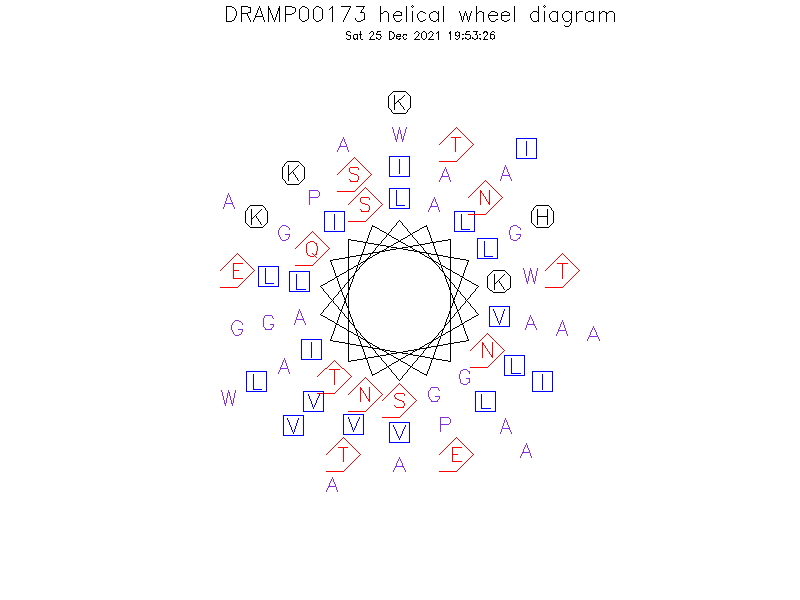
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP00173.
Physicochemical Information
-
Formula
- C282H460N74O77
Absent Amino Acids
- CDFMRY
Common Amino Acids
- A
Mass
- 6119.2
PI
- 9.53
Basic Residues
- 5
Acidic Residues
- 2
Hydrophobic Residues
- 35
Net Charge
- +3
-
Boman Index
- 35.71
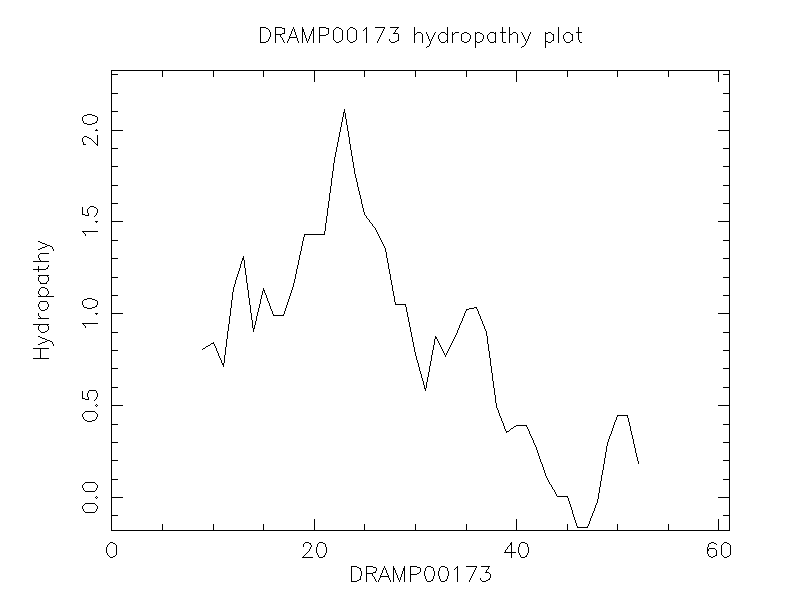
Hydrophobicity
- 0.751
Aliphatic Index
- 129.84
Half Life
-
- Mammalian:5.5 hour
- Yeast:3 min
- E.coli:2 min
Extinction Coefficient Cystines
- 16500
Absorbance 280nm
- 275
Polar Residues
- 16
DRAMP00173
Comments Information
Function
- This precursor showed high similarity to the precursor of lactocyclicin Q, a cyclic bacteriocin produced by Lactococcus sp. strain QU 12. This is the first report of a cyclic bacteriocin produced by a strain of the genus Leuconostoc.
Literature Information
- ·Literature 1
-
Title
- Identification and characterization of leucocyclicin Q, a novel cyclic bacteriocin produced by Leuconostoc mesenteroides TK41401.
-
Pubmed ID
- 21948835
-
Reference
- Appl Environ Microbiol. 2011 Nov;77(22):8164-8170.
-
Author
- Masuda Y, Ono H, Kitagawa H, Ito H, Mu F, Sawa N, Zendo T, Sonomoto K.