General Information
-
DRAMP ID
- DRAMP00017
-
Peptide Name
- Microbisporicin A1 (Bacteriocin)
-
Source
- Microbispora corallina (Gram-positive bacteria)
-
Family
- Belongs to the lantibiotic family (Class I bacteriocin)
-
Gene
- Not found
-
Sequence
- VTSWSLCTPGCTSPGGGSNCSFCC
-
Sequence Length
- 24
-
UniProt Entry
- No entry found
-
Protein Existence
- Protein level
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial
-
Target Organism
-
- Human pathogens:
Human Pathogen Activity L100 Staphylococcus aureus ATCC6538P MIC≤0.13 μg/ml L819 Staphylococcus aureus Smith ATCC19636 MIC≤0.13 μg/ml L1400 S. aureus MRSA MIC≤0.13 μg/ml L613 S. aureus MRSA MIC≤0.13 μg/ml L3798 S. aureus VISA MIC=2 μg/ml L3798 Staphylococcus epidermidis ATCC12228 MIC≤0.13 μg/ml L1729 Staphylococcus haemolyticus met-r MIC=8 μg/ml L49 Streptococcus pyogenes MIC≤0.13 μg/ml L44 Streptococcus pneumoniae MIC≤0.13 μg/ml L559 Enterococcus faecalis MIC=1 μg/ml L560 Enterococcus faecalis Van A MIC=0.5 μg/ml LA533 Enterococcus faecalis Van A MIC=1 μg/ml L568 Enterococcus faecium MIC=2 μg/ml L569 Enterococcus faecium Van A MIC=1 μg/ml LB518 E. faecium Van A MIC=2 μg/ml L884 Lactobacillus garviae MIC=1 μg/ml L148 Lactobacillus delbrueckii ATCC04797 MIC=4 μg/ml L3607 Clostridium perfringens ATCC13124 MIC≤0.125 μg/ml L4018 Clostridium difficile MIC≤0.125 μg/ml L4043 Clostridium butyricum MIC≤0.125 μg/ml Propionibacterium granulosum ATCC25564 MIC=0.03 μg/ml L1329 Propionibacterium acnes MIC=0.5 μg/ml Propionibacterium limphophylum ATCC27250 MIC=0.015 μg/ml L970 Haemophilus influenzae ATCC19418 MIC=32 μg/ml L76 Moraxella catarrhalis ATCC8176 MIC=0.25 μg/ml L1613 Neisseria meningitidis ATCC13090V MIC=0.5 μg/ml L997 Neisseria gonorrhoeae MIC=0.25 μg/ml
- Human pathogens:
-
Hemolytic Activity
-
- No hemolysis information or data found in the reference(s) presented in this entry
-
Cytotoxicity
- No cytotoxicity information found in the reference(s) presented
-
Binding Target
- Not found
Structure Information
-
Linear/Cyclic
- Cyclic
-
N-terminal Modification
- Free
-
C-terminal Modification
- Amidation and Cyclization
-
Nonterminal Modifications and Unusual Amino Acids
- ①There are five thioether intramolecular bridges which link Ala3 and Ala7, Ala8 and Ala11, Ala13 and Ala20, Ala18 and Ala23, respectively. ②The residue at position 2 is (Z)-2,3-didehydrobutyrine (Dhb). ③The residue at position 4 is chloro-tryptophan (ClTrp). ④The residue at position 5 is 2,3-didehydroalanine. ⑤The residue at position 8 is 2-Aminobutyric acid (Abu). ⑥The residue at position 14 is bis-hydroxylated proline.
-
Stereochemistry
- L
-
Structure
- Rich
-
Structure Description
- Not found
-
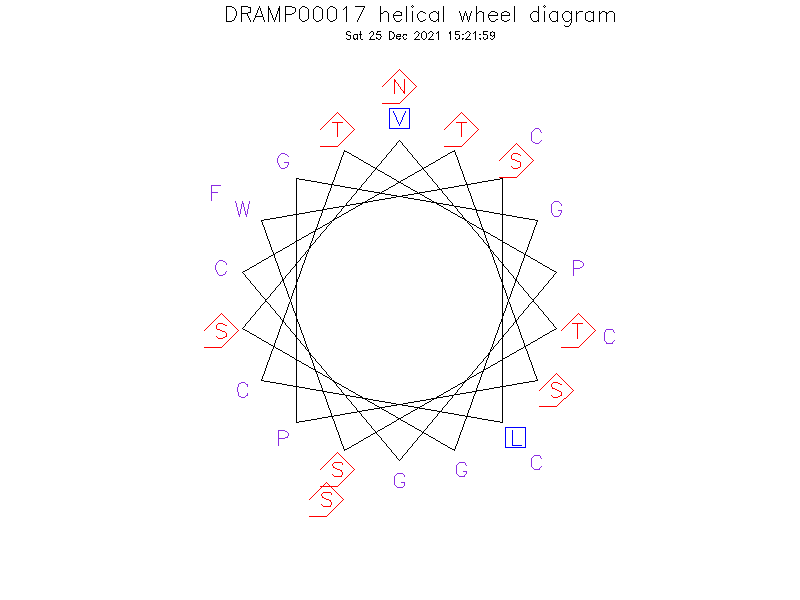
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP00017.
Physicochemical Information
-
Formula
- C95H144N26O34S5
Absent Amino Acids
- ADEHIKMQRY
Common Amino Acids
- CS
Mass
- 2354.64
PI
- 5.47
Basic Residues
- 0
Acidic Residues
- 0
Hydrophobic Residues
- 4
Net Charge
- 0
-
Boman Index
- -6.92
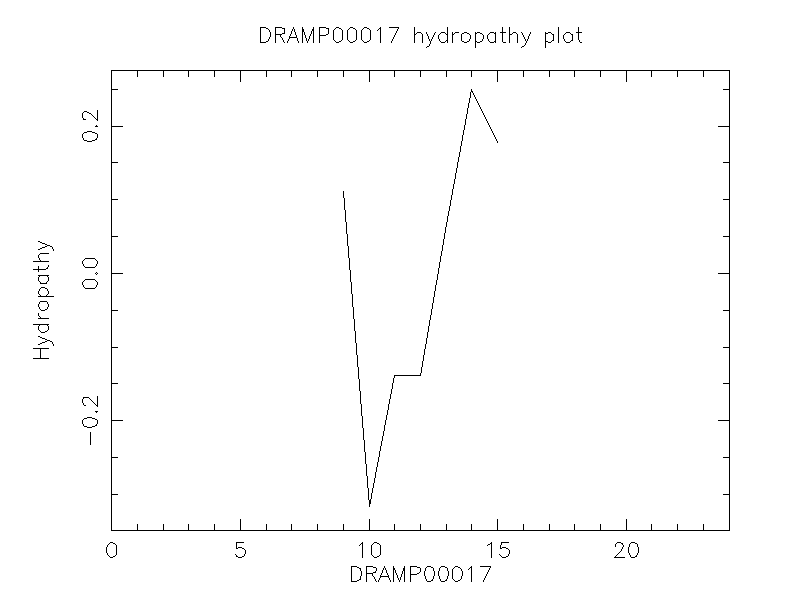
Hydrophobicity
- 0.333
Aliphatic Index
- 28.33
Half Life
-
- Mammalian:100 hour
- Yeast:>20 hour
- E.coli:>10 hour
Extinction Coefficient Cystines
- 5750
Absorbance 280nm
- 250
Polar Residues
- 18
DRAMP00017
Comments Information
Function
- Microbisporicins A1 displayed similar antibacterial activity by selectively blocking peptidoglycan biosynthesis, leading to cytoplasmic accumulation of the UDP-linked precursor.
PTM
- Microbisporicin A1 contains five ether rings
Literature Information
- ·Literature 1
-
Title
- Determining the structure and mode of action of microbisporicin, a potent lantibiotic active against multiresistant pathogens.
-
Pubmed ID
- 18215770
-
Reference
- Chem Biol. 2008 Jan;15(1):22-31.
-
Author
- Castiglione F, Lazzarini A, Carrano L, Corti E, Ciciliato I, Gastaldo L, Candiani P, Losi D, Marinelli F, Selva E, Parenti F.