General Information
-
DRAMP ID
- DRAMP00039
-
Peptide Name
- Pep5 (Bacteriocin)
-
Source
- Staphylococcus epidermidis (Gram-positive bacteria)
-
Family
- Belongs to the type A lantibiotic family (Class I bacteriocin)
-
Gene
- pepA
-
Sequence
- TAGPAIRASVKQCQKTLKATRLFTVSCKGKNGCK
-
Sequence Length
- 34
-
UniProt Entry
- P19578
-
Protein Existence
- Protein level
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial
-
Target Organism
- No MICs found in DRAMP database
-
Hemolytic Activity
-
- No hemolysis information or data found in the reference(s) presented in this entry
-
Cytotoxicity
-
- Not included yet
-
Binding Target
- Lipid II
Structure Information
-
Linear/Cyclic
- Not included yet
-
N-terminal Modification
- Not included yet
-
C-terminal Modification
- Not included yet
-
Nonterminal Modifications and Unusual Amino Acids
- Not included yet
-
Stereochemistry
- Not included yet
-
Structure
- Rich
-
Structure Description
- Not found
-
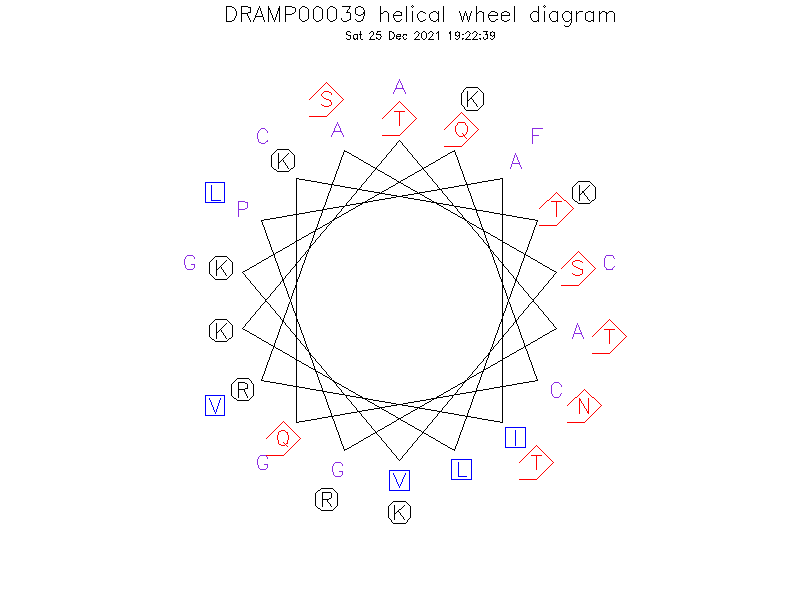
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP00039.
Physicochemical Information
-
Formula
- C153H269N49O44S3
Absent Amino Acids
- DEHMWY
Common Amino Acids
- K
Mass
- 3595.3
PI
- 10.33
Basic Residues
- 8
Acidic Residues
- 0
Hydrophobic Residues
- 10
Net Charge
- +8
-
Boman Index
- -58.22
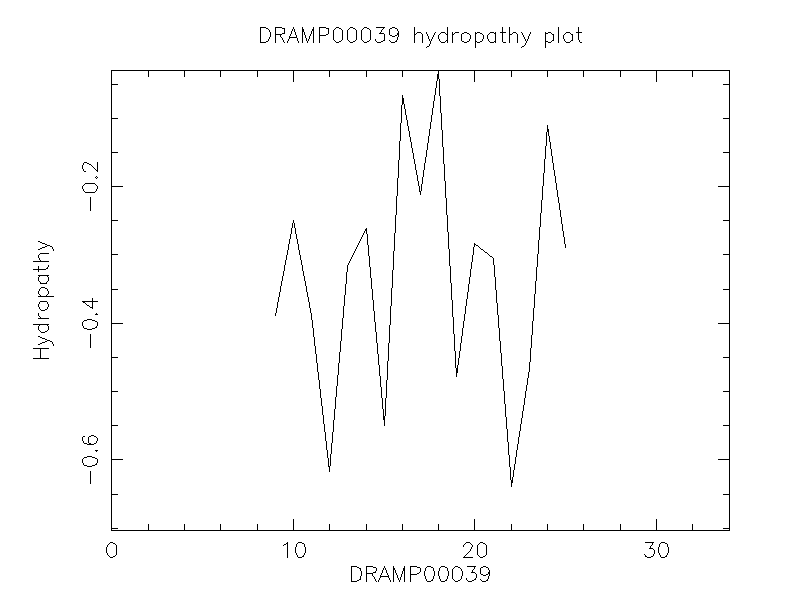
Hydrophobicity
- -0.356
Aliphatic Index
- 63.24
Half Life
-
- Mammalian:7.2 hour
- Yeast:>20 hour
- E.coli:>10 hour
Extinction Coefficient Cystines
- 125
Absorbance 280nm
- 3.79
Polar Residues
- 13
DRAMP00039
Comments Information
Function
- Lanthionine-containing peptide antibiotic (lantibiotic) active on Gram-positive bacteria. The bactericidal activity of lantibiotics is based on depolarization of energized bacterial cytoplasmic membranes, initiated by the formation of aqueous transmembrane pores.
PTM
- Maturation of lantibiotics involves the enzymic conversion of Thr, and Ser into dehydrated AA and the formation of thioether bonds with cysteine. This is followed by membrane translocation and cleavage of the modified precursor.After proteolysis of the propeptide, the N-terminal 2,3-didehydrobutyrine hydrolyzes to 2-oxobutanoic acid, possibly spontaneously.
Literature Information
- ·Literature 1
-
Title
- Pep5, a new lantibiotic: structural gene isolation and prepeptide sequence.
-
Pubmed ID
- 2764678
-
Reference
- Arch Microbiol. 1989;152(1):16-19.
-
Author
- Kaletta C, Entian KD, Kellner R, Jung G, Reis M, Sahl HG.
- ·Literature 2
-
Title
- Nucleotide sequence of the lantibiotic Pep5 biosynthetic gene cluster and functional analysis of PepP and PepC. Evidence for a role of PepC in thioether formation.
-
Pubmed ID
- 7556197
-
Reference
- Eur J Biochem. 1995 Sep 1;232(2):478-489.
-
Author
- Meyer C, Bierbaum G, Heidrich C, Reis M, S¼ling J, Iglesias-Wind MI, Kempter C, Molitor E, Sahl HG.