General Information
-
DRAMP ID
- DRAMP00066
-
Peptide Name
- Lacticin Q (Bacteriocin)
-
Source
- Lactococcus lactis QU 5 (Gram-positive bacteria)
-
Family
- Belongs to the class II bacteriocin
-
Gene
- lnqQ
-
Sequence
- AGFLKVVQLLAKYGSKAVQWAWANKGKILDWLNAGQAIDWVVSKIKQILGIK
-
Sequence Length
- 52
-
UniProt Entry
- A4UVR2
-
Protein Existence
- Predicted
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial, Anti-Gram+
-
Target Organism
-
- Gram-positive bacteria:
Target Organism Activity Bacillus cereus JCM 2152 - Bacillus circulans JCM 2504 - Bacillus coagulans JCM 2257 - Bacillus subtilis JCM 1465 - Enterococcus faecalis JCM 5803 - Enterococcus faecium JCM 5804 - Enterococcus mundtii QU 2 - Enterococcus hirae ATCC 10541 - Lactococcus lactis ATCC 19435 - Lactococcus lactis NCDO 497 - Lactobacillus acidophilus JCM 1132 - Lactobacillus alimentarius JCM 1095 - Lactobacillus brevis JCM 1059 - Lactobacillus casei JCM 1134 - Lactobacillus coryniformis JCM 1164 - Lactobacillus sakeii JCM 1157 - Leuconostoc mesenteroides JCM 6124 - Listeria innocua ATCC 33090 - Micrococcus luteus IFO 12708 - Pediococcus pentosaceus JCM 5885 -
- Gram-positive bacteria:
-
Hemolytic Activity
-
- No hemolysis information or data found in the reference(s) presented in this entry
-
Cytotoxicity
-
- Not included yet
-
Binding Target
- Not found
Structure Information
-
Linear/Cyclic
- Not included yet
-
N-terminal Modification
- Not included yet
-
C-terminal Modification
- Not included yet
-
Nonterminal Modifications and Unusual Amino Acids
- Not included yet
-
Stereochemistry
- Not included yet
-
Structure
- Not found
-
Structure Description
- Not found
-
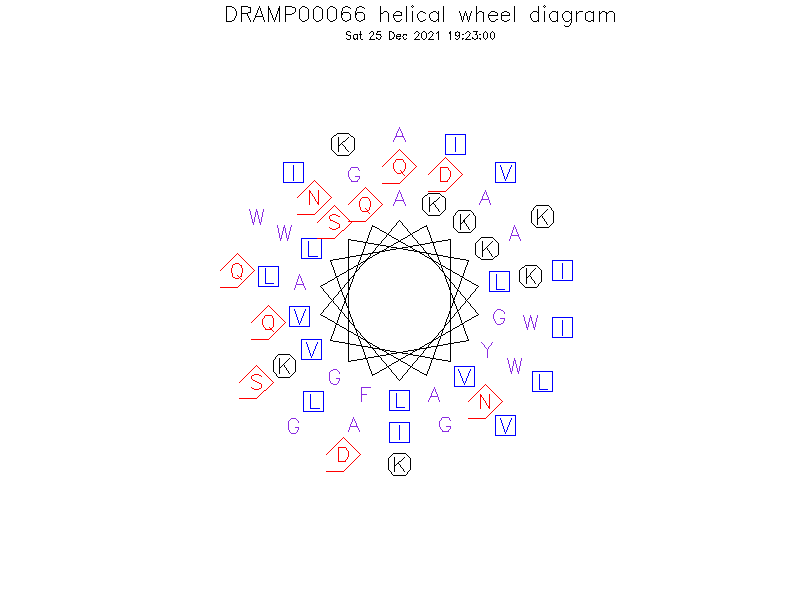
Helical Wheel Diagram
-
Predicted Structure
- There is no predicted structure for DRAMP00066.
Physicochemical Information
-
Formula
- C274H436N70O66
Absent Amino Acids
- CEHMPRT
Common Amino Acids
- KA
Mass
- 5766.9
PI
- 10.1
Basic Residues
- 8
Acidic Residues
- 2
Hydrophobic Residues
- 28
Net Charge
- +6
-
Boman Index
- -0.23
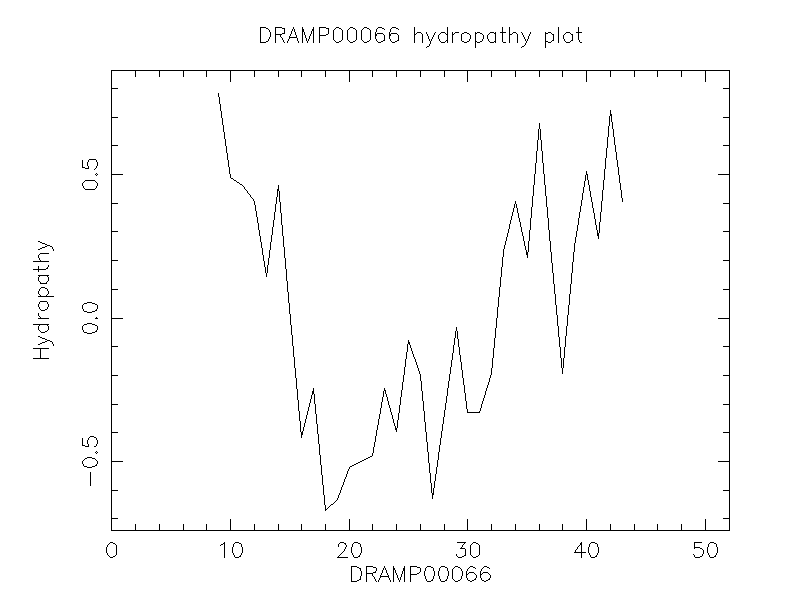
Hydrophobicity
- 0.269
Aliphatic Index
- 123.85
Half Life
-
- Mammalian:4.4 hour
- Yeast:>20 hour
- E.coli:>10 hour
Extinction Coefficient Cystines
- 23490
Absorbance 280nm
- 460.59
Polar Residues
- 10
DRAMP00066
Comments Information
Lacticin Q was very stable against heat treatment and changes in pH; in particular, it was stable at alkaline pH values, while nisin A was inactivated. Moreover, lacticin Q induced ATP efflux from a Listeria sp. strain in a shorter time and at a lower concentration than nisin A, indicating that the former affected indicator cells in a different manner from that of the latter.
Literature Information
- ·Literature 1
-
Title
- Structural analysis and characterization of lacticin Q, a novel bacteriocin belonging to a new family of unmodified bacteriocins of gram-positive bacteria.
-
Pubmed ID
- 17351096
-
Reference
- Appl Environ Microbiol. 2007 May;73(9):2871-2877.
-
Author
- Fujita K, Ichimasa S, Zendo T, Koga S, Yoneyama F, Nakayama J, Sonomoto K.