General Information
-
DRAMP ID
- DRAMP00373
-
Peptide Name
- Seed non-specific lipid transfer protein-like (ns-LTP; Plants)
-
Source
- Raphanus sativus (Radish)
-
Family
- Belongs to the plant LTP family
-
Gene
- Not found
-
Sequence
- ALSCGTVNSNLAACIGYLTQNAPLARGCCTGVTNLNNMAXTTP
-
Sequence Length
- 43
-
UniProt Entry
- P29420
-
Protein Existence
- Protein level
Activity Information
-
Biological Activity
- Antimicrobial, Antifungal
-
Target Organism
-
- Fungi: Alternaria brassicola (MUCL 20297) (IC50=48 µg/mL), Ascochyta pisi (MUCL 20164) (IC50=41 µg/mL), Botrytis cinerea (MUCL 30158) (IC50=45 µg/mL), Colletotrichum lindemuthianum (MUCL 9577) (IC50=25 µg/mL), Fusarium culmorum (IMI 180420) (IC50=20 µg/mL), Fusarium oxysporum f. sp. lycopersici (MUCL 909) (IC50=54 µg/mL), Fusarium oxysporum f. sp. pisi (IMI 236441) (IC50=58 µg/mL), Nectria haematococca (Collection Van Etten 26022) (IC50=100 µg/mL), Phoma betae (MUCL 9916) (IC50=18 µg/mL), Pyricularia oryzae (MUCL 30166) (IC50=10 µg/mL), Trichoderma hamatum (MUCL 29736) (IC50=30 µg/mL), Verticillium dahliae (MUCL 6963) (IC50=7 µg/mL).
-
Hemolytic Activity
-
- No hemolysis information or data found in the reference(s) presented in this entry
-
Cytotoxicity
-
- Not included yet
-
Binding Target
- Lipid-binding
Structure Information
-
Linear/Cyclic
- Not included yet
-
N-terminal Modification
- Not included yet
-
C-terminal Modification
- Not included yet
-
Nonterminal Modifications and Unusual Amino Acids
- Not included yet
-
Stereochemistry
- Not included yet
-
Structure
- Not found
-
Structure Description
- Not found
-
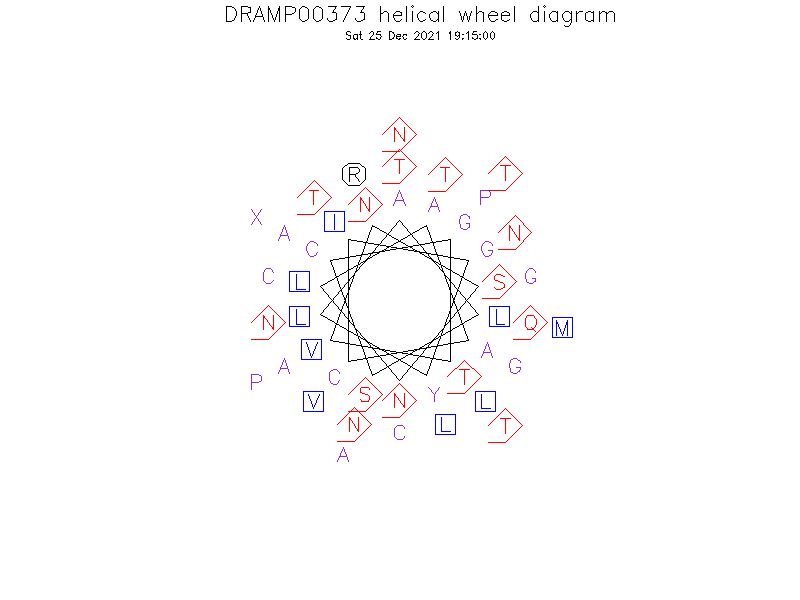
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP00373.
Physicochemical Information
-
Formula
- C173H286N52O58S5
Absent Amino Acids
- DEFHKW
Common Amino Acids
- ANT
Mass
- 4312.13
PI
- 7.92
Basic Residues
- 1
Acidic Residues
- 0
Hydrophobic Residues
- 14
Net Charge
- +1
-
Boman Index
- -22.97
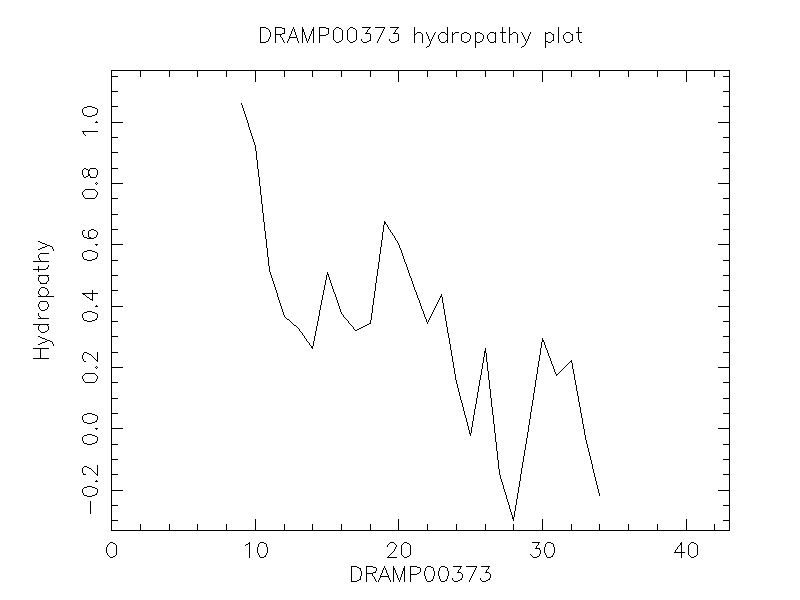
Hydrophobicity
- 0.319
Aliphatic Index
- 81.86
Half Life
-
- Mammalian:4.4 hour
- Yeast:>20 hour
- E.coli:>10 hour
Extinction Coefficient Cystines
- 1740
Absorbance 280nm
- 41.43
Polar Residues
- 23
DRAMP00373
Comments Information
Function
- Plant non-specific lipid-transfer proteins transfer phospholipids as well as galactolipids across membranes. May play a role in wax or cutin deposition in the cell walls of expanding epidermal cells and certain secretory tissues. This isoform inhibits the hyphal growth of several fungi in vitro.
Literature Information
- ·Literature 1
-
Title
- In vitro antifungal activity of a radish (Raphanus sativus L.) seed protein homologous to non-specific lipid transfer proteins.
-
Pubmed ID
- 16653017
-
Reference
- Plant Physiol. 1992 Oct;100(2):1055-1058.
-
Author
- Terras FR, Goderis IJ, Van Leuven F, Vanderleyden J, Cammue BP, Broekaert WF.