General Information
-
DRAMP ID
- DRAMP02934
-
Peptide Name
- PM1-S (linear derivative of PM1)
-
Source
- Synthetic construct
-
Family
- Not found
-
Gene
- Not found
-
Sequence
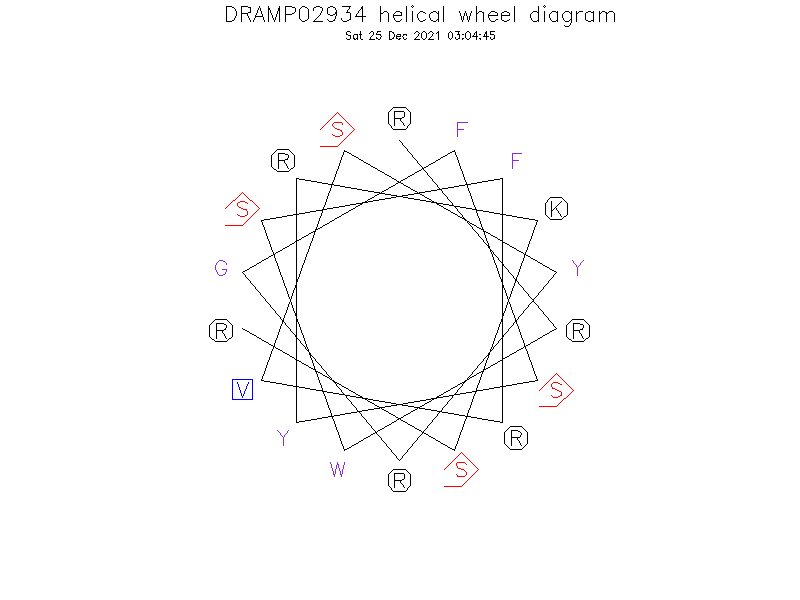
- RRWSFRVSYRGFSYRKSR
-
Sequence Length
- 18
-
UniProt Entry
- No entry found
-
Protein Existence
- Synthetic
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial, Anti-Gram+, Anti-Gram-, Antifungal
-
Target Organism
-
- Gram-negative bacteria:
Target Organism Activity Escherichia coli UB1005 MIC=4 µg/ml E. coli DC2 MIC=4 µg/ml Salmonella typhimurium (defensin-sensitive) MIC=2 µg/ml S. typhimurium MIC=8 µg/ml Pseudomonas aeruginosa K799 MIC=32 µg/ml P. aeruginosa Z61 (MIC=16 µg/ml); - - Gram-positive bacteria:
Target Organism Activity Staphylococcus aureus SAP0017 MIC=32 µg/ml Staphylococcus epidermidis MIC=16 µg/ml Enterococcus faecalis MIC=2 µg/ml - Yeasy: Candida albicans (MIC=64 µg/ml).
- Gram-negative bacteria:
-
Hemolytic Activity
-
- No hemolysis information or data found in the reference(s) presented in this entry
-
Cytotoxicity
-
- Not included yet
-
Binding Target
- Not found
Structure Information
-
Linear/Cyclic
- Not included yet
-
N-terminal Modification
- Not included yet
-
C-terminal Modification
- Not included yet
-
Nonterminal Modifications and Unusual Amino Acids
- Not included yet
-
Stereochemistry
- Not included yet
-
Structure
- No secondary structure (CD)
-
Structure Description
- The CD spectra of PM1-S indicates that, in all environments (phosphate buffer, TFE, and liposomes), the peptide displays no observable patterns that could be related to structural features. This indicates that PM1-S is a flexible molecule with no favoured conformation.
-
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP02934.
Physicochemical Information
-
Formula
- C108H164N38O25
Absent Amino Acids
- ACDEHILMNPQT
Common Amino Acids
- R
Mass
- 2394.73
PI
- 12.01
Basic Residues
- 7
Acidic Residues
- 0
Hydrophobic Residues
- 4
Net Charge
- +7
-
Boman Index
- -95.68
Hydrophobicity
- -1.567
Aliphatic Index
- 16.11
Half Life
-
- Mammalian:1 hour
- Yeast:2 min
- E.coli:2 min
Extinction Coefficient Cystines
- 8480
Absorbance 280nm
- 498.82
Polar Residues
- 7
DRAMP02934
Comments Information
Function
- PM1-S remaines active against Gram-negative bacteria but displayed poor activity to both Gram-positive and fungal microorganisms.
Literature Information
- ·Literature 1
-
Title
- Structure-activity relationships for the beta-hairpin cationic antimicrobial peptide polyphemusin I.
-
Pubmed ID
- 15134657
-
Reference
- Biochim Biophys Acta. 2004 May 6;1698(2):239-250.
-
Author
- Powers JP, Rozek A, Hancock RE.