General Information
-
DRAMP ID
- DRAMP18276
-
Peptide Name
- HalR1(Bacteriocin)
-
Source
- Halobacterium salinarum GN101
-
Family
- Not found
-
Gene
- Not found
-
Sequence
- LQSNININTAAAVILIFNQVQVGALCAPTPVSGGGPPP
-
Sequence Length
- 38
-
UniProt Entry
- No entry found
-
Protein Existence
Activity Information
-
Biological Activity
- Archaeolytic
-
Target Organism
- No MICs found in DRAMP database
-
Hemolytic Activity
-
- No hemolysis information or data found in the reference(s) presented in this entry
-
Cytotoxicity
-
- Not included yet
-
Binding Target
- Not found
Structure Information
-
Linear/Cyclic
- Not included yet
-
N-terminal Modification
- Not included yet
-
C-terminal Modification
- Not included yet
-
Nonterminal Modifications and Unusual Amino Acids
- Not included yet
-
Stereochemistry
- Not included yet
-
Structure
- Not found
-
Structure Description
- HalR1 is 63% identical and 71% similar to HalS8.
-
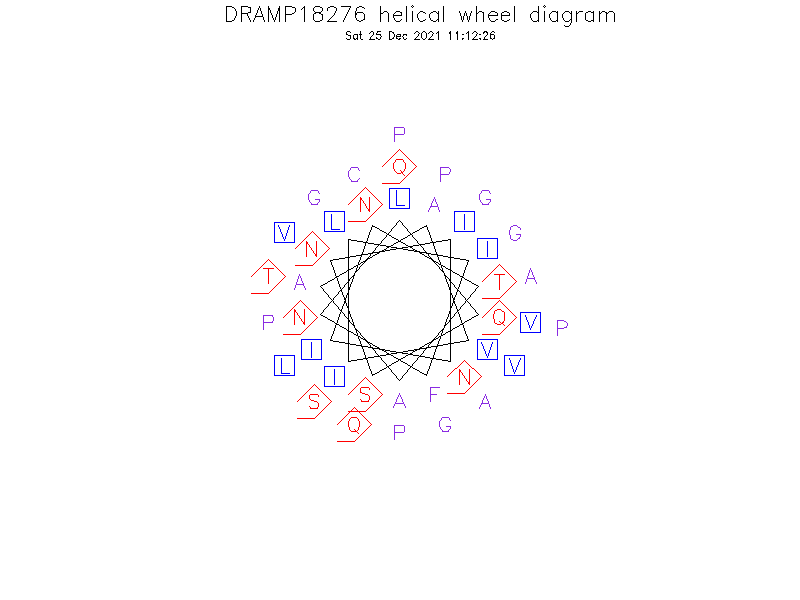
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP18276.
Physicochemical Information
-
Formula
- C167H273N45O50S
Absent Amino Acids
- DEHKMRWY
Common Amino Acids
- AP
Mass
- 3743.34
PI
- 5.52
Basic Residues
- 0
Acidic Residues
- 0
Hydrophobic Residues
- 17
Net Charge
- 0
-
Boman Index
- 1255
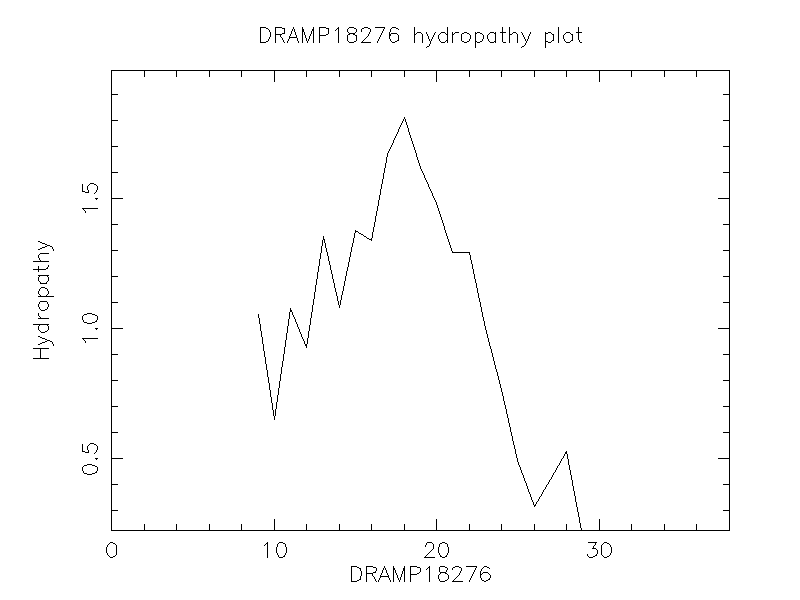
Hydrophobicity
- 0.616
Aliphatic Index
- 115.53
Half Life
-
- Mammalian:5.5 hour
- Yeast:3 min
- E.coli:2 min
Extinction Coefficient Cystines
- 0
Absorbance 280nm
- 0
Polar Residues
- 13
DRAMP18276
Comments Information
Unlike HalH6, HalR1 does not appear to be archaeolytic, since it causes no change in the optical density or cell morphology of stationary phase Hbt. salinarum, nor can it form zones of inhibition on fully grown lawns. HalR1 has a dose-dependent, archaeostatic effect on growing Hbt. salinarum.
Literature Information
- ·Literature 1
-
Title
- Halocins and sulfolobicins: the emerging story of archaeal protein and peptide antibiotics.
-
Pubmed ID
- 11938468
-
Reference
- J Ind Microbiol Biotechnol. 2002 Jan;28(1):23-31.
-
Author
- O'Connor EM, Shand RF.