General Information
-
DRAMP ID
- DRAMP18583
-
Peptide Name
- Tricystine cyclic cystine TP (ccTP, Tachyplesin-1 peptide derivative)
-
Source
- Synthetic construct
-
Family
- Derived from the peptide Tachyplesin-1
-
Gene
- Not found
-
Sequence
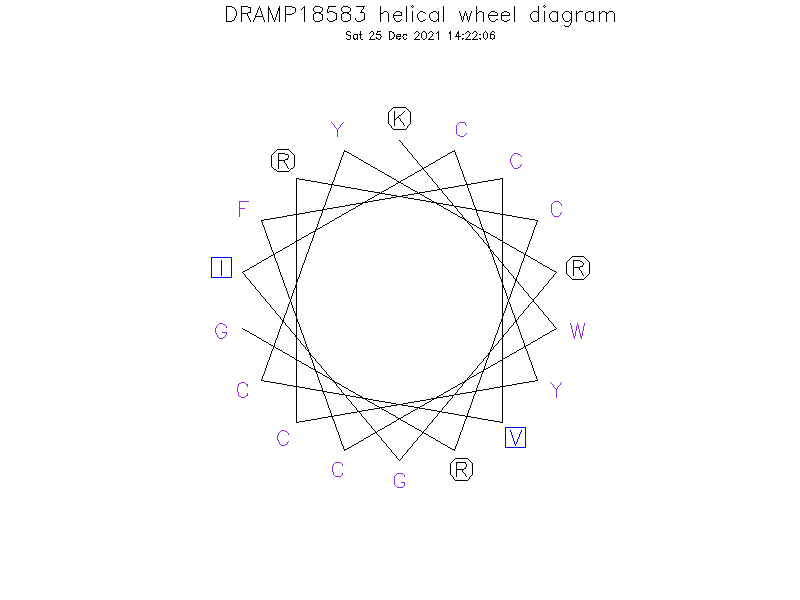
- KWCFCVCYRGICYCRCRG
-
Sequence Length
- 18
-
UniProt Entry
- No entry found
-
Protein Existence
- Synthetic
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial, Anti-Gram+, Anti-Gram-, Anitifungal
-
Target Organism
-
- [Ref.10673369]Gram-positive bacteria: Staphylococcus aureus (MIC=1.0 μM in low-salt; MIC=0.8 μM in high-salt), Micrococcus luteus (MIC=0.5 μM in low-salt; MIC=0.4 μM in high-salt), Enterococcus faecalis (MIC=1.0 μM in low-salt; MIC=0.9 μM in high-salt);
- Gram-negative bacteria: Escherichia coli (MIC=3.0 μM in low-salt; MIC=6.9 μM in high-salt), Pseudomonas aeruginosa (MIC=5.0 μM in low-salt; MIC=5.2 μM in high-salt), Primula vulgaris (MIC=6.0 μM in low-salt; MIC=14.4 μM in high-salt), Klebsiella oxytoca (MIC=2.1 μM in low-salt; MIC=7.8 μM in high-salt);
- Fungi: Candida albicans (MIC=5.1 μM in low-salt; MIC=17.2 μM in high-salt), Candida kefyr (MIC=3.9 μM in low-salt; MIC=4.1 μM in high-salt), Candida tropicalis (MIC=5.2 μM in low-salt; MIC=1.1 μM in high-salt).
-
Hemolytic Activity
-
- [Ref.10673369] EC25=3800 μM against human red blood cells
-
Cytotoxicity
- No cytotoxicity information found in the reference(s) presented
-
Binding Target
- Not found
Structure Information
-
Linear/Cyclic
- Cyclic
-
N-terminal Modification
- No specific N-terminal
-
C-terminal Modification
- No specific C-terminal
-
Nonterminal Modifications and Unusual Amino Acids
- Disulfide bonds between Cys3 and Cys16,Cys5 and Cys14,Cys7 and Cys12.
-
Stereochemistry
- L
-
Structure
- Beta tile
-
Structure Description
- Although there is insufficient information to deconvolute these CD spectra of these two-b-strand cyclic cystine-knot structures due to the various contributions by β-turns, aromatic residues, disulfides and b-sheet structures, it is reasonable to conclude that all cyclic tachyplesins 2–6 as well as RTD display ordered structures in methanol.
-
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP18583.
Physicochemical Information
-
Formula
- C95H143N29O21S6
Absent Amino Acids
- ADEHLMNPQST
Common Amino Acids
- C
Mass
- 2219.72
PI
- 8.94
Basic Residues
- 4
Acidic Residues
- 0
Hydrophobic Residues
- 4
Net Charge
- +4
-
Boman Index
- -2676
Hydrophobicity
- 0.267
Aliphatic Index
- 37.78
Half Life
-
- Mammalian:1.3 hour
- Yeast:3 min
- E.coli:2 min
Extinction Coefficient Cystines
- 8855
Absorbance 280nm
- 520.88
Polar Residues
- 10
DRAMP18583
Comments Information
Function
- Antimicrobial activity against Gram-positive bacteria, Gram-negative bacteria.Antifungal activity agaist Candida albicans, Candida kefyr, Candida tropicalis
Literature Information
- ·Literature 1
-
Title
- Marked increase in membranolytic selectivity of novel cyclic tachyplesins constrained with an antiparallel two-beta strand cystine knot framework.
-
Pubmed ID
- 10673369
-
Reference
- Biochem Biophys Res Commun. 2000 Jan 27;267(3):783-90.
-
Author
- Tam JP, Lu YA, Yang JL.