General Information
-
DRAMP ID
- DRAMP20768
-
Peptide Name
- Pore-forming peptide ameobapore C (EH-APP; Parasite, amoebozoa, protozoa, protists)
-
Source
- Entamoeba histolytica
-
Family
- Belongs to the amoebapore (amoebapore) family
-
Gene
- Not found
-
Sequence
- IPVLCPVCTSLVGKLIDLVLGGAVDKVTDYLETLCAKADGLVETLCTKIVSYGIDKLIEKILEGGSAKLICGLIHAC
-
Sequence Length
- 77
-
UniProt Entry
- Q24825
-
Protein Existence
- Protein level
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial, Anti-Gram+, Antibiotic
-
Target Organism
-
- [Ref.7715451] Gram-positive bacterium: Micrococcus luteus (MIC=5 μM, MBC=10 μM)
-
Hemolytic Activity
-
- No hemolysis information or data found in the reference(s) presented in this entry
-
Cytotoxicity
-
- Not included yet
-
Binding Target
- Not found
Structure Information
-
Linear/Cyclic
- Not included yet
-
N-terminal Modification
- Not included yet
-
C-terminal Modification
- Not included yet
-
Nonterminal Modifications and Unusual Amino Acids
- Disulfide bonds between Cys5 and Cys77, Cys8 and Cys71, Cys35 and Cys46
-
Stereochemistry
- Not included yet
-
Structure
- Alpha helix
-
Structure Description
- Not found
-
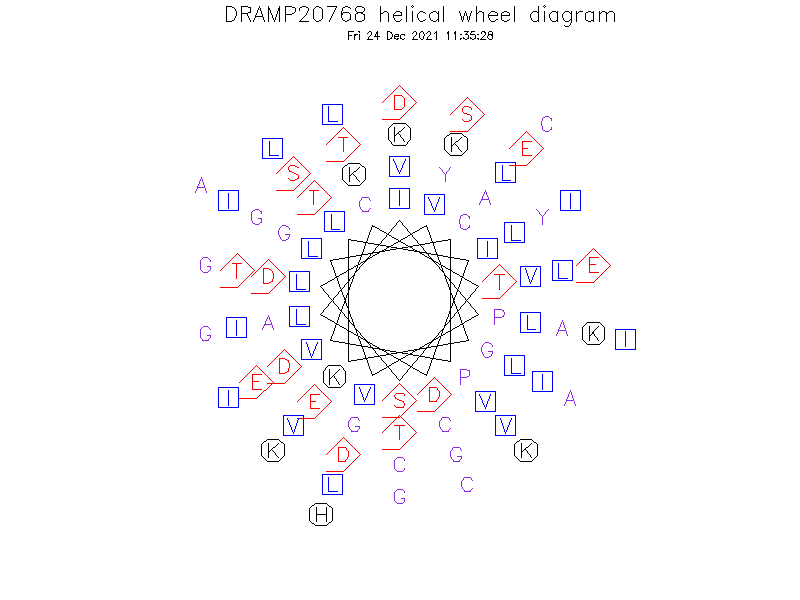
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP20768.
Physicochemical Information
-
Formula
- C360H610N86O106S6
Absent Amino Acids
- FMNQRW
Common Amino Acids
- L
Mass
- 8031.68
PI
- 5.12
Basic Residues
- 8
Acidic Residues
- 9
Hydrophobic Residues
- 34
Net Charge
- -1
-
Boman Index
- 2221
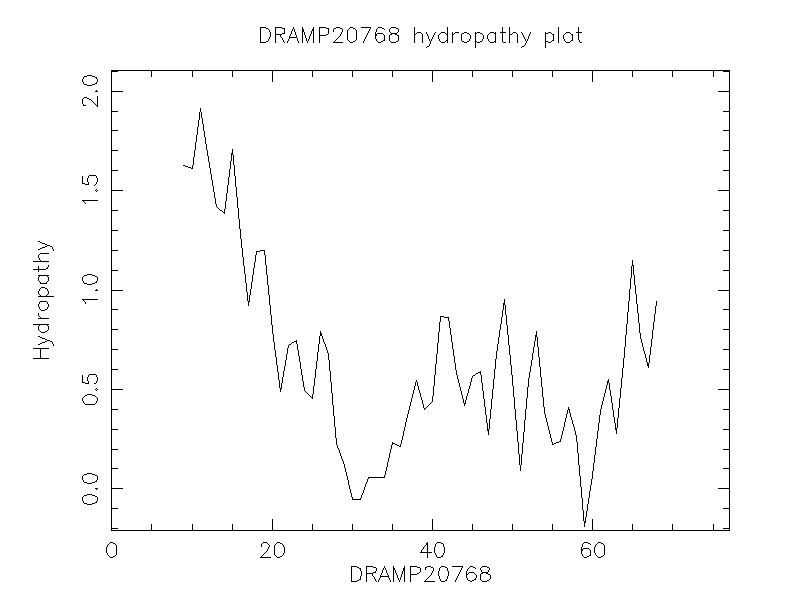
Hydrophobicity
- 0.858
Aliphatic Index
- 142.99
Half Life
-
- Mammalian:20 hour
- Yeast:30 min
- E.coli:>10 hour
Extinction Coefficient Cystines
- 3355
Absorbance 280nm
- 44.14
Polar Residues
- 24
DRAMP20768
Comments Information
Function
- Forms pores in the cytoplasmic membrane of host cells. Has antibacterial activity against M.luteus, no activity against E.coli. Implicated in the cytolytic activity of the parasite.
Literature Information
- ·Literature 1
-
Title
- Amoebapores, a family of membranolytic peptides from cytoplasmic granules of Entamoeba histolytica: isolation, primary structure, and pore formation in bacterial cytoplasmic membranes.
-
Pubmed ID
- 7715451
-
Reference
- Mol Microbiol. 1994 Dec;14(5):895-904.
-
Author
- Leippe M, Andrä J, Nickel R, Tannich E, Müller-Eberhard HJ.