General Information
-
DRAMP ID
- DRAMP20966
-
Peptide Name
- andricin B (Andrias davidianus, Amphibians, Animals)
-
Source
- Andrias davidianus
-
Family
- Not found
-
Gene
- Not found
-
Sequence
- GLTRLFSVIK
-
Sequence Length
- 10
-
UniProt Entry
- No entry found
-
Protein Existence
- Not found
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial, Anti-Gram+, Anti-Gram-, Antifungal
-
Target Organism
-
- [Ref.29130500]Gram-positive bacteria : Bacillus subtilis CICC 10034 (MIC=32 μg/ml, 33.4 μMol/ml;MBC=32 μg/ml, 33.4 μMol/ml), Staphylococcus aureus CICC10306 (MIC=16 μg/ml, 16.7 μMol/ml;MBC=32 μg/ml, 33.4 μMol/ml), Staphylococcus aureus CICC 10384 (MIC=16 μg/ml, 16.7 μMol/ml;MBC=32 μg/ml, 33.4 μMol/ml), Listeria innocua CICC 10417 (MIC=32 μg/ml, 33.4 μMol/ml;MBC=32 μg/ml, 33.4 μMol/ml);
- Gram-negative bacteria : Escherichia coli CICC10293 (MIC=16 μg/ml, 16.7 μMol/ml;MBC=32 μg/ml, 33.4 μMol/ml), Pseudomonas aeruginosa CICC 21636 (MIC=8 μg/ml, 8.3 μMol/ml;MBC=16 μg/ml, 16.7 μMol/ml), Serratia marcescens ATCC 4112 (MIC=8 μg/ml, 8.3 μMol/ml;MBC=16 μg/ml, 16.7 μMol/ml), Enterobacter cloacae CICC 21539 (MIC=16 μg/ml, 16.7 μMol/ml;MBC=32 μg/ml, 33.4 μMol/ml), Salmonella paratyphi CICC 10437 (MIC=32 μg/ml, 33.4 μMol/ml;MBC=32 μg/ml, 33.4 μMol/ml);
- Fungi : Aspergillus niger CICC 2124 (MIC=64 μg/ml, 66.8 μMol/ml;MBC=64 μg/ml, 66.8 μMol/ml), Candida albicans CICC 1965 (MIC=32 μg/ml, 33.4 μMol/ml;MBC=64 μg/ml, 66.8 μMol/ml), Saccharomyces cerevisiae CICC 1002 (MIC=64 μg/ml, 66.8 μMol/ml;MBC=64 μg/ml, 66.8 μMol/ml)
-
Hemolytic Activity
-
- [Ref.29130500] 0% hemolysis at 64 μg/ml , 0.5% hemolysis at 128 μg/ml , 2% hemolysis at 256 μg/ml , 13% hemolysis at 512 μg/ml against human red blood cells
-
Cytotoxicity
- No cytotoxicity information found in the reference(s) presented
-
Binding Target
- Not found
Structure Information
-
Linear/Cyclic
- Linear
-
N-terminal Modification
- Free
-
C-terminal Modification
- Free
-
Nonterminal Modifications and Unusual Amino Acids
- None
-
Stereochemistry
- L
-
Structure
- Random coil
-
Structure Description
- The residues Lys and Arg possess the positive charge, and charge-induced folding may influence the structure.
-
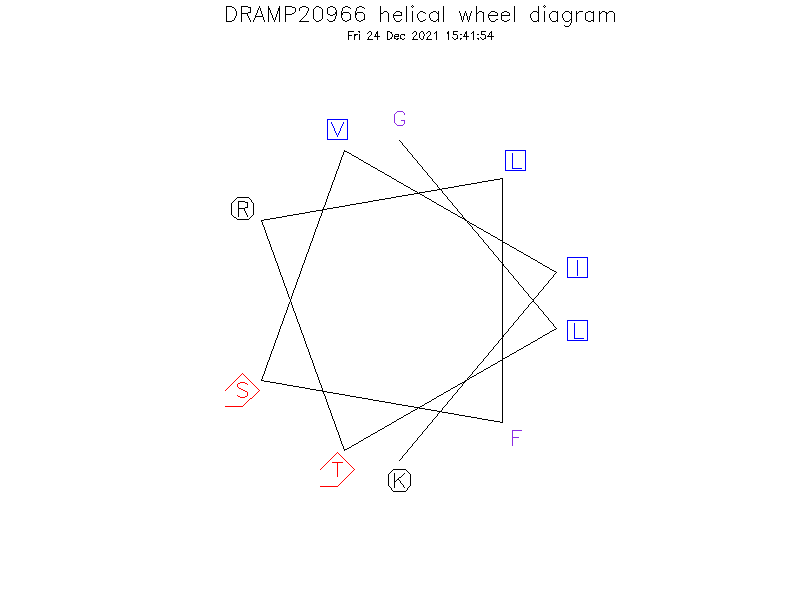
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP20966.
Physicochemical Information
-
Formula
- C53H92N14O13
Absent Amino Acids
- ACDEHMNPQWY
Common Amino Acids
- L
Mass
- 1133.4
PI
- 11
Basic Residues
- 2
Acidic Residues
- 0
Hydrophobic Residues
- 5
Net Charge
- +2
-
Boman Index
- -372
Hydrophobicity
- 0.88
Aliphatic Index
- 146
Half Life
-
- Mammalian:30 hour
- Yeast:>20 hour
- E.coli:>10 hour
Extinction Coefficient Cystines
- 0
Absorbance 280nm
- 0
Polar Residues
- 3
DRAMP20966
Comments Information
Function
- Antibacterial activity against Gram-positive bacteria and Gram-negative bacteria, Antifungal activity agaist Candida albicans, Saccharomyces cerevisiae, Aspergillus niger
Literature Information
- ·Literature 1
-
Title
- Purifification, characterization and application of a novel antimicrobial peptide from Andrias davidianus blood
-
Pubmed ID
- 29130500
-
Reference
- Lett Appl Microbiol.?2018 Jan;66(1):38-43. doi: 10.1111/lam.12823.
-
Author
- J. Pei, Z. Feng,T. Ren,H. Sun,H. Han,W. Jin,J. Dang and Y. Tao