General Information
-
DRAMP ID
- DRAMP21006
-
Peptide Name
- [G1K, K8R]cGm (Derived from Gm)
-
Source
- Synthetic construct
-
Family
- Not found
-
Gene
- Not found
-
Sequence
- KCRRLCYRQRCVTYCRGR
-
Sequence Length
- 18
-
UniProt Entry
- No entry found
-
Protein Existence
- Synthetic form
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial, Anti-Gram+, Anti-Gram-, Antifungal, Antitumor
-
Target Organism
-
- [Ref.28741926]Gram-positive bacteria : Staphylococcuss aureus ATCC 25923 (MIC=2 μM);
- Gram-negative bacteria : Escherichia coli ATCC 25922 (MIC=0.5-1 μM), Escherichia coli CGSC 5167 (MIC<0.015 μM), Klebsiella pneumoniae ATCC 700603 (MIC=8 μM), Acinetobacter baumannii ATCC 19606 (MIC=1-2 μM), Pseudomonas aeruginosa ATCC 27853 (MIC<0.25 μM), Helicobacter pylori ATCC 43504 (MIC>80 μM);
- Fungi : Candida albicans ATCC 90028 (MIC=2 μM), Cryptococcus neoformans ATCC 208821 (MIC=0.125-0.25 μM)
-
Hemolytic Activity
-
- [Ref.28741926] 0% hemolysis at 0.5 μM , 2.5% hemolysis at 0.6 μM , 20% hemolysis at 8 μM , 43.3-56.9% hemolysis at 64 μM , 50% hemolysis at 100 μM against human red blood cells
-
Cytotoxicity
-
- [Ref.28741926] CC50 = 19.5 ± 0.5 μM against CRL-1739, CC50 = 1.7 ± 0.1 μM against MM96L, CC50 = 9.0 ± 0.5 μM against HFF-1, CC50 = 44.2 ± 2.2μM against HeLa, CC50 = 6.7 ± 0.7 μM against MCF-7, CC50 = 2.1 ± 0.2 μM against K-562, CC50 = 9.0 ± 1.1μM against HL-60;CC50=5.1 ± 0.3μM against PBMCs.
-
Binding Target
- Not found
Structure Information
-
Linear/Cyclic
- Cyclic
-
N-terminal Modification
- No specific N-terminal
-
C-terminal Modification
- No specific C-terminal
-
Nonterminal Modifications and Unusual Amino Acids
- Disulfide bonds between Cys2 and Cys15, Cys6 and Cys11.
-
Stereochemistry
- L
-
Structure
- Not found
-
Structure Description
- Not found
-
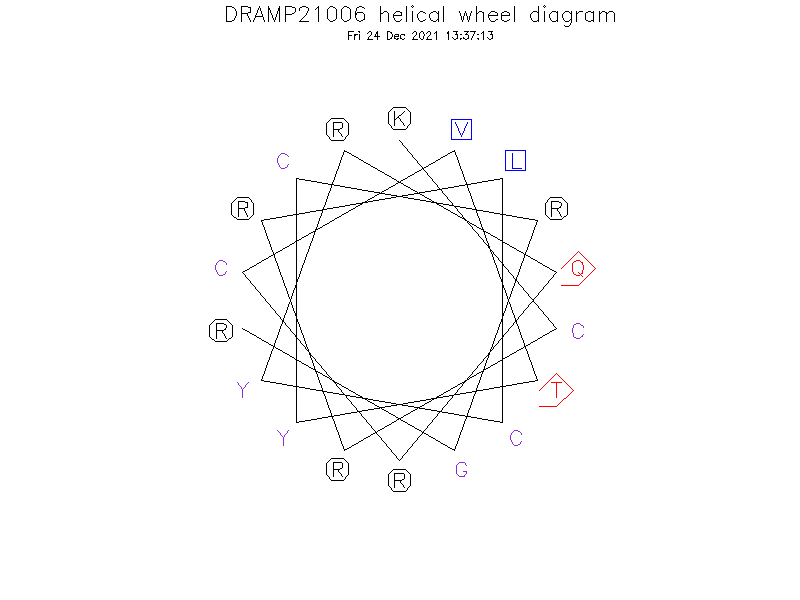
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP21006.
Physicochemical Information
-
Formula
- C94H162N38O23S4
Absent Amino Acids
- ADEFHIMNPSW
Common Amino Acids
- R
Mass
- 2320.8
PI
- 10.33
Basic Residues
- 7
Acidic Residues
- 0
Hydrophobic Residues
- 2
Net Charge
- +7
-
Boman Index
- -8844
Hydrophobicity
- -1.117
Aliphatic Index
- 37.78
Half Life
-
- Mammalian:1.3 hour
- Yeast:3 min
- E.coli:2 min
Extinction Coefficient Cystines
- 3230
Absorbance 280nm
- 190
Polar Residues
- 8
DRAMP21006
Comments Information
Function
- Antibacterial activity against Gram-positive bacteria and Gram-negative bacteria, Antifungal activity agaist Candida albicans, Cryptococcus neoformans
Literature Information
- ·Literature 1
-
Title
- Redesigned Spider Peptide with Improved Antimicrobial and Anticancer Properties
-
Pubmed ID
- 28741926
-
Reference
- ACS Chem Biol. 2017 Sep 15;12(9):2324-2334. doi: 10.1021/acschembio.7b00459.?
-
Author
- Sonia Troeira Henriques, Nicole Lawrence, Stephanie Chaousis, Anjaneya S. Ravipati, Olivier Cheneval,Aurelie H. Benfield, Alysha G. Elliott, Angela Maria Kavanagh, Matthew A. Cooper, Lai Yue Chan,Yen-Hua Huang, and David J. Craik