General Information
-
DRAMP ID
- DRAMP21085
-
Peptide Name
- PRW4 (PR) (Derived from PMAP-36)
-
Source
- Synthetic construct
-
Family
- Not found
-
Gene
- Not found
-
Sequence
- RFRRLRWKTRWRLKKI
-
Sequence Length
- 16
-
UniProt Entry
- No entry found
-
Protein Existence
- Synthetic form
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial, Anti-Gram+, Anti-Gram-
-
Target Organism
-
- [Ref.25818457] Gram-positive bacteria : Staphylococcus aureus 25923(MIC=73.6 mg/L), Staphylococcus aureus 29213(MIC=9.2 mg/L), Staphylococcus epidermidis 12228(MIC=18.4 mg/L), Enterococcus faecalis 29212(MIC=9.2 mg/L), Bacillus subtilis 11060(MIC=18.4 mg/L);
- Gram-negative bacteria : Escherichia coli 078(MIC=9.2 mg/L), Escherichia coli K88(MIC=9.2 mg/L), Escherichia coli ER2566(MIC=18.4 mg/L), Escherichia coli 44102(MIC=18.4 mg/L), Escherichia coli 1515(MIC=9.2 mg/L), Escherichia coli 25922(MIC=9.2 mg/L), Escherichia coli UB1005(MIC=9.2 mg/L), Salmonella typhimurium 7731(MIC=18.4 mg/L), Salmonella typhimurium 1402(MIC=18.4 mg/L), Pseudomonas aeruginosa 21625(MIC=73.6 mg/L), Pseudomonas aeruginosa 21630(MIC=36.8 mg/L), Pseudomonas aeruginosa 10419(MIC=36.8 mg/L), Pseudomonas aeruginosa 27853(MIC>294.3 mg/L), Salmonella pullorum 7913(MIC>294.3 mg/L)
-
Hemolytic Activity
-
- [Ref.25818457] 0 % hemolysis at 294.3 mg/L against human red blood cells
-
Cytotoxicity
-
- [Ref.25818457] The cell viability of PRW4 is 100%, 96%, 92%, 86%, 85%, 83%, 82%, 80% and 80% at peptide concentrations of 1.1, 2.3, 4.6, 9.2, 18.4, 36.8, 73.6, 147.1 and 294.3 μg/ml
-
Binding Target
- Not found
Structure Information
-
Linear/Cyclic
- Linear
-
N-terminal Modification
- Free
-
C-terminal Modification
- Amidation
-
Nonterminal Modifications and Unusual Amino Acids
- None
-
Stereochemistry
- L
-
Structure
- Random coil, alpha helix, beta Sheet
-
Structure Description
- Not found
-
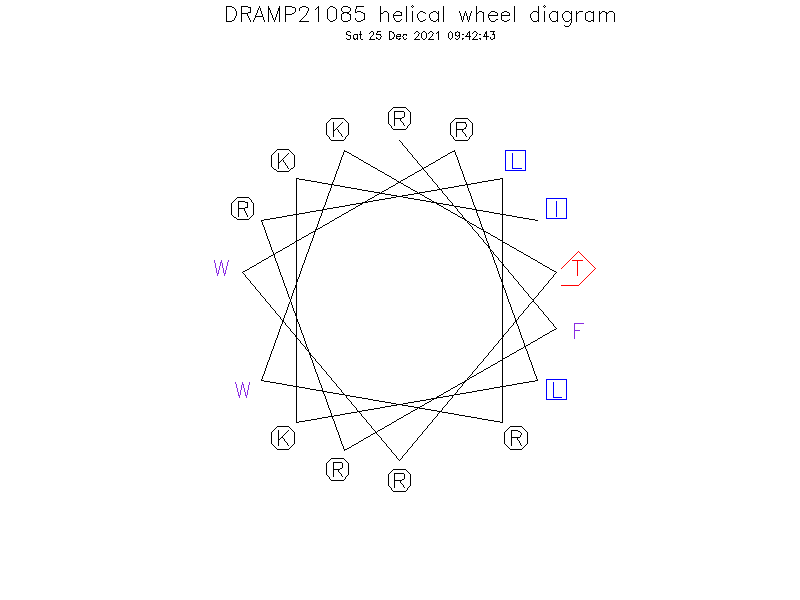
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP21085.
Physicochemical Information
-
Formula
- C107H179N39O18
Absent Amino Acids
- ACDEGHMNPQSVY
Common Amino Acids
- R
Mass
- 2299.85
PI
- 12.7
Basic Residues
- 9
Acidic Residues
- 0
Hydrophobic Residues
- 6
Net Charge
- +9
-
Boman Index
- -8634
Hydrophobicity
- -1.644
Aliphatic Index
- 73.13
Half Life
-
- Mammalian:1 hour
- Yeast:2 min
- E.coli:2 min
Extinction Coefficient Cystines
- 11000
Absorbance 280nm
- 733.33
Polar Residues
- 1
DRAMP21085
Comments Information
Function
- Antibacterial activity against Gram-positive bacteria and Gram-negative bacteria
Literature Information
- ·Literature 1
-
Title
- Characterization of cell selectivity, physiological stability and endotoxin neutralization capabilities of a-helix-based peptide amphiphiles
-
Pubmed ID
- 25818457
-
Reference
- Biomaterials. 2015 Jun;52:517-30. doi: 10.1016/j.biomaterials.2015.02.063.
-
Author
- Zhi Ma, Dandan Wei, Ping Yan, Xin Zhu, Anshan Shan , Zhongpeng Bi