General Information
-
DRAMP ID
- DRAMP21416
-
Peptide Name
- WV (De Novo Synthesis)
-
Source
- synthetic construct
-
Family
- Not found
-
Gene
- Not found
-
Sequence
- WVKAAAKAAAKVW
-
Sequence Length
- 13
-
UniProt Entry
- No entry found
-
Protein Existence
- Synthetic form
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial, Anti-Gram+, Anti-Gram-
-
Target Organism
-
- [Ref.30897850] Gram-positive bacteria:Staphylococcus aureus ATCC 27853(MIC>128μM), Staphylococcus epidermidis ATCC 12228(MIC>128μM), Staphylococcus faecalis ATCC 29212(MIC=64μM), Bacillus subtilis CMCC 63501(MIC>128μM);
- Gram-negative bacteria:Escherichia coli ATCC 25922 (MIC=32μM), Escherichia coli UB 1005 (MIC>128μM), Pseudomonas aeruginosa ATCC 27853(MIC>128μM), Salmonella typhimurium ATCC 14028(MIC>128μM)
-
Hemolytic Activity
-
- [Ref.30897850] 1.5% hemolysis at 256μM against human red blood cells
-
Cytotoxicity
-
- [Ref.30897850] ①The cell viability of RAW 264.7 induced by WV is 102.5%, 93.5%, 97.8%, 95.7%, 99.3%, 99.6%, 93.1% at peptide concentrations of 1, 2, 4, 8, 16, 32, 64 μM, while the cytotoxicity of WV on RAW 264.7 cells (Control samples) is 93.1%, 67.8%, 37.7%, 17.8%, 0%, 1.4% and 1.1% at peptide concentrations of 1, 2, 4, 8, 16, 32, 64 μM.②The cell viability of HEK293T induced by WV is 115.2%, 104.3%, 114.5%, 107.6%, 105.8%, 108.0% and 93.8% at peptide concentrations of 1, 2, 4, 8, 16, 32, 64 μM, while the cytotoxicity of WV on HEK293T cells (Control samples) is 101.4%, 80.1%, 31.9%, 1.4%, 1.4%, 1.8% and 1.1% at peptide concentrations of 1, 2, 4, 8, 16, 32, 64 μM.
-
Binding Target
- Not found
Structure Information
-
Linear/Cyclic
- Linear
-
N-terminal Modification
- Free
-
C-terminal Modification
- Amidaton
-
Nonterminal Modifications and Unusual Amino Acids
- None
-
Stereochemistry
- L
-
Structure
- α-helix predicted by I-TASSER, random coil in PBS, α-helix in SDS
-
Structure Description
- Not found
-
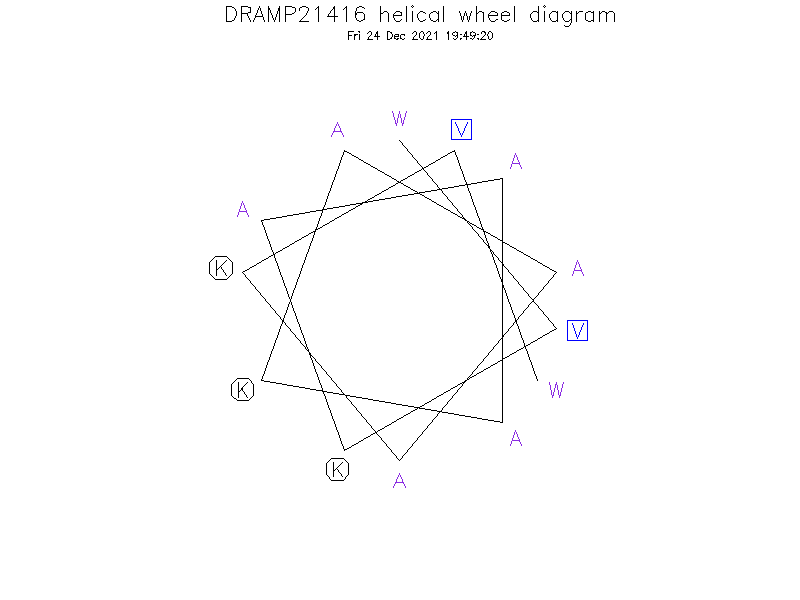
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP21416.
Physicochemical Information
-
Formula
- C68H106N18O14
Absent Amino Acids
- CDEFGHILMNPQRSTY
Common Amino Acids
- A
Mass
- 1399.7
PI
- 10.3
Basic Residues
- 3
Acidic Residues
- 0
Hydrophobic Residues
- 10
Net Charge
- +3
-
Boman Index
- 695
Hydrophobicity
- 0.439
Aliphatic Index
- 90.77
Half Life
-
- Mammalian:2.8 hour
- Yeast:3 min
- E.coli:2 min
Extinction Coefficient Cystines
- 11000
Absorbance 280nm
- 916.67
Polar Residues
- 0
DRAMP21416
Comments Information
Function
- Antibacterial activity against Gram-positive bacteria and Gram-nagetive bacteria.
Literature Information
- ·Literature 1
-
Title
- Symmetrical Modification of Minimized Dermaseptins to Extend the Spectrum of Antimicrobials with Endotoxin Neutralization Potency
-
Pubmed ID
- 30897850
-
Reference
- Int J Mol Sci. 2019 Mar; 20(6): 1417. Published online 2019 Mar 20. doi: 10.3390/ijms20061417
-
Author
- Changxuan Shao, Weizhong Li, Peng Tan, Anshan Shan, Xiujing Dou, Deying Ma, Chunyu Liu