General Information
-
DRAMP ID
- DRAMP29048
-
Peptide Name
- PA-13
-
Source
- Synthetic construct
-
Family
- Not found
-
Gene
- N/A
-
Sequence
- KIAKRIWKILRRR
-
Sequence Length
- 13
-
UniProt Entry
- No entry found
-
Protein Existence
- Synthetic form
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial, Anti-Gram+, Anti-Gram-
-
Target Organism
-
- [Ref.32499514] Gram-negative bacteria: Pseudomonas aeruginosa ATCC 27853 (MIC = 3.91 µg/ml), Escherichia coli ATCC 25922 (MIC = 31.25 µg/ml), Escherichia coli O157:H7 MT strain (MIC = 31.25 µg/ml), Shigella sonnei ATCC 11060 (MIC = 3.91 µg/ml), Salmonella Typhimurium ATCC 13311 (MIC = 1.95 µg/ml), Vibrio cholera O1 Inaba DMST 16261 (MIC = 31.25 µg/ml), Acinetobacter baumannii MT strain (MIC = 7.81 µg/ml).
- Clinically isolated MDR P. aeruginosa: P. aeruginosa No.1 (3.91 μg/mL), P. aeruginosa No.2 (7.81μg/mL), P. aeruginosa No.3 (7.81 μg/mL), P. aeruginosa No.4 (7.81 μg/mL), P. aeruginosa No.5 (7.81 μg/mL), P. aeruginosa No.6 (7.81 μg/mL), P. aeruginosa No.7 (7.81 μg/mL), P. aeruginosa No.8 (7.81 μg/mL), P. aeruginosa No.9 (7.81 μg/mL), P. aeruginosa No.10 (7.81 μg/mL), P. aeruginosa No.11 (7.81 μg/mL), P. aeruginosa No.12 (3.91 μg/mL), P. aeruginosa No.13 (15.63 μg/mL), P. aeruginosa No.14 (7.81 μg/mL),
- Gram-positive bacteria:Staphylococcus aureus ATCC 25923 (MIC = 62.5 µg/ml) Staphylococcus epidermidis ATCC 12228 (MIC = 3.91 µg/ml), Bacillus cereus ATCC 11778 (MIC = 15.63 µg/ml), Enterococcus faecalis ATCC 29212 (MIC = 250 µg/ml), Listeria monocytogenes 10403 s (MIC = 3.91 µg/ml)
-
Hemolytic Activity
-
- [Ref.32499514]MHC = 125 μg/mL. Note:MHC is the minimum concentration that caused 10% hemolysis of human red blood cells (hRBC). When no 10% hemolysis was observed at 250 µg/ml, a value of 500 µg/ml was used to calculate the therapeutic index.
-
Cytotoxicity
-
- [Ref.32499514]Cytotoxic activity of PA-13 against the L929 mouse fibroblast cell line was determined by a colorimetric MTT viability assay. At the MIC (3.91 µg/ml), the cell survival was up to 90%. Even at the maximum concentration (250 µg/ml), the survival was common (81.29%). Melittin, used as a positive control, showed a strong and dose-dependent cytotoxic effect; no living cells were detected at 62.5 µg/ml. Peptide PA-13 displayed very little cytotoxicity on L929 cells when compared to melittin.
-
Binding Target
- Membrane
Structure Information
-
Linear/Cyclic
- Linear
-
N-terminal Modification
- Free
-
C-terminal Modification
- Amidation
-
Nonterminal Modifications and Unusual Amino Acids
- None
-
Stereochemistry
- L
-
Structure
- α-helical
-
Structure Description
- Not found
-
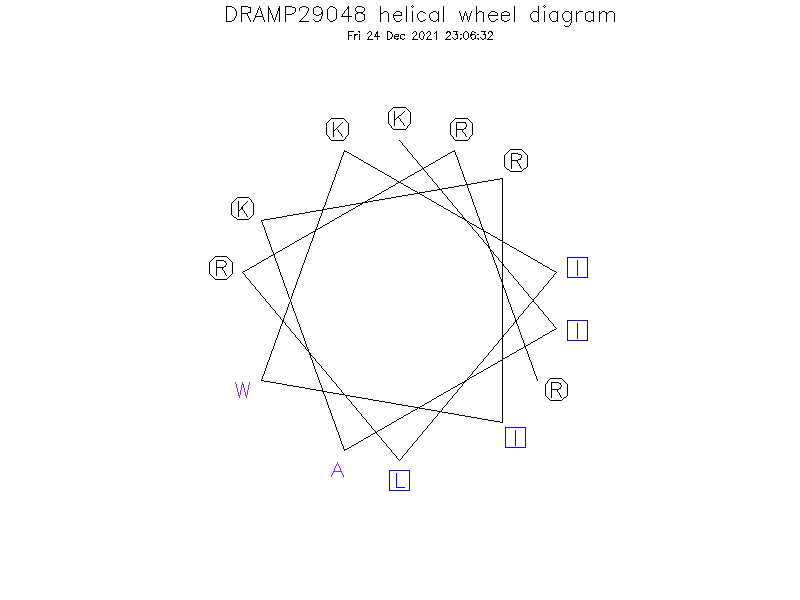
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP29048.
Physicochemical Information
-
Formula
- C80H145N29O14
Absent Amino Acids
- CDEFGHMNPQSTVY
Common Amino Acids
- R
Mass
- 1737.22
PI
- 12.48
Basic Residues
- 7
Acidic Residues
- 0
Hydrophobic Residues
- 6
Net Charge
- +7
-
Boman Index
- -5251
Hydrophobicity
- -0.885
Aliphatic Index
- 127.69
Half Life
-
- Mammalian:1.3 hour
- Yeast:3 min
- E.coli:2 min
Extinction Coefficient Cystines
- 5500
Absorbance 280nm
- 458.33
Polar Residues
- 0
DRAMP29048
Comments Information
Function
- PA-13 exhibited potent antibacterial activity against P. aeruginosa, including multidrug resistant strains, via penetration through the bacteria’s membrane and causing depolarization, permeabilization and rupture of membranes, and subsequent leakage of intracellular contents and cell death. The peptide displayed low hemolytic and cytotoxic activities against mammalian cells suggesting a therapeutic potential. Furthermore, PA-13 showed anti-inflammatory activity via LPS neutralization. Note
Literature Information
- ·Literature 1
-
Title
- A novel, rationally designed, hybrid antimicrobial peptide, inspired by cathelicidin and aurein, exhibits membrane-active mechanisms against Pseudomonas aeruginosa
-
Pubmed ID
- 32499514
-
Reference
- Sci Rep. 2020 Jun 4;10(1):9117. doi: 10.1038/s41598-020-65688-5.
-
Author
- Klubthawee N, Adisakwattana P, Hanpithakpong W, Somsri S, Aunpad R