General Information
-
DRAMP ID
- DRAMP29076
-
Peptide Name
- dCATH 16 (modified from dCATH)
-
Source
- Synthetic construct
-
Family
- Not found
-
Gene
- N/A
-
Sequence
- QLVPLAIKIYRAWKRR
-
Sequence Length
- 16
-
UniProt Entry
- No entry found
-
Protein Existence
- Synthetic form
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial, Anti-Gram-, Anti-Gram+
-
Target Organism
-
- [Ref.32698035] Gram-positive bacteria: S. aureus KCTC 1621 (MIC = 8 μM), S. epidermidis KCTC 1917 (MIC = 16 μM), B. subtilis KCTC 3068 (MIC = 16 μM);
- Resistant Gram-positive bacteria: MRSA CCARM 3089 (MIC = 32 μM), MRSA CCARM 3090 (MIC = 16 μM), MRSA CCARM 3095 (MIC = 32 μM), VREF ATCC 51559(MIC = 32 μM);
- Gram-negative bacteria: E. coli KCTC 1682 (MIC = 32 μM), P. aeruginosa KCTC 1637(MIC = 16 μM), S. typhimurium KCTC 1926 (MIC = 16 μM);
- Resistant Gram-negative bacteria: MDRPAd CCARM 2095 (MIC = 32 μM), MDRPA CCARM 2109 (MIC = 32 μM);
- Yeast: C. albicans KCTC 7965 (MIC = 8 μM), C. albicans KCTC 7121 (MIC = 16 μM).
-
Hemolytic Activity
-
- [Ref.32698035] HC10 = 180 μM. Note: HC10 is the peptide concentration that caused 10% hemolysis of sheep red blood cells (sRBCs).
-
Cytotoxicity
-
- [Ref.32698035] At the concentration of 1.25, 2.5, 5, 10, 20, 40, 80 μM, the cell survival of dCATH 16 against human bone marrow SH-SY5Y cells is 98%, 95%, 91%, 86%, 54%, 36%, 32%.
-
Binding Target
- Not found
Structure Information
-
Linear/Cyclic
- Linear
-
N-terminal Modification
- Free
-
C-terminal Modification
- Amidation
-
Nonterminal Modifications and Unusual Amino Acids
- None
-
Stereochemistry
- L
-
Structure
- Unknown
-
Structure Description
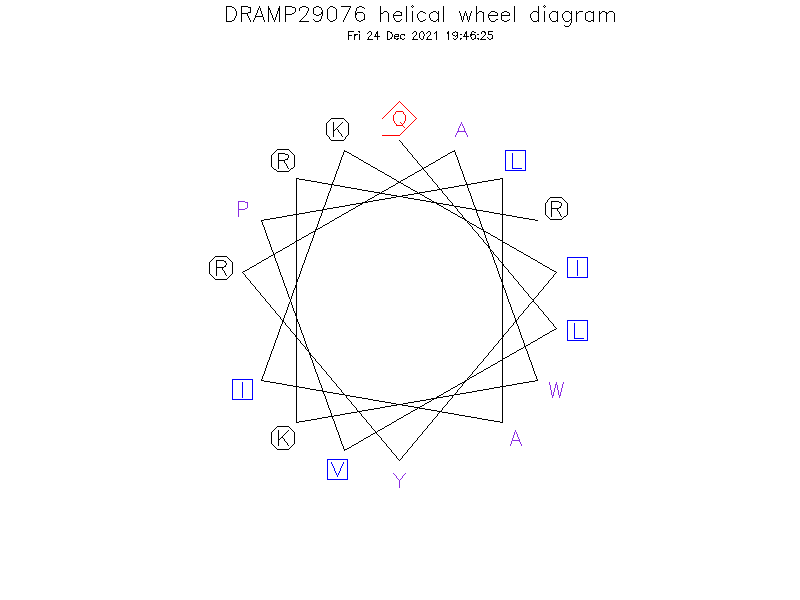
- Initially, four analog peptides were designed by truncating two, four, six, and eight amino acids at the N-terminal ends of dCATH peptide. dCATH and its truncated analogs could not adopt the amphipathic α-helical structure since the hydrophobic and charged residues were dispersed all around the molecule.
-
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP29076.
Physicochemical Information
-
Formula
- C95H159N29O19
Absent Amino Acids
- CDEFGHMNST
Common Amino Acids
- R
Mass
- 2011.49
PI
- 11.73
Basic Residues
- 5
Acidic Residues
- 0
Hydrophobic Residues
- 8
Net Charge
- +5
-
Boman Index
- -3187
Hydrophobicity
- -0.263
Aliphatic Index
- 128.13
Half Life
-
- Mammalian:0.8 hour
- Yeast:10 min
- E.coli:>10 hour
Extinction Coefficient Cystines
- 6990
Absorbance 280nm
- 466
Polar Residues
- 1
DRAMP29076
Comments Information
Comment
- The peptide induces potent antimicrobial activity.
Literature Information
- ·Literature 1
-
Title
- Antimicrobial and anti-inflammatory activities of short dodecapeptides derived from duck cathelicidin: Plausible mechanism of bactericidal action and endotoxin neutralization
-
Pubmed ID
- 32698035
-
Reference
- Eur J Med Chem. 2020 Oct 15;204:112580. doi: 10.1016/j.ejmech.2020.112580. Epub 2020 Jul 16.
-
Author
- Kumar SD, Shin SY