General Information
-
DRAMP ID
- DRAMP29092
-
Peptide Name
- Hn-Mc (a chimeric peptide comprised of the N-terminus of HPA3NT3 and the C-terminus of melittin)
-
Source
- Synthetic construct
-
Family
- Not found
-
Gene
- N/A
-
Sequence
- FKRLKKLISWIKRKRQQ
-
Sequence Length
- 17
-
UniProt Entry
- No entry found
-
Protein Existence
- Synthetic construct
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial, Anti-Gram+, Anti-Gram-, Antifungal, Anti-inflammatory
-
Target Organism
-
- [Ref.32731574]Yeast: C. albicans (MIC = 16 μM), C. krusei (MIC = 8 μM), C. parapsilosis (MIC = 16 μM), C. tropicalis (MIC = 4 μM), T. beigellii (MIC = 1 μM);
- Mold: T. rubrum (MIC = 1-2 μM), F. moniliforme(MIC = 1-2 μM), F. solani(MIC = 1 μM), F. oxysporum(MIC = 1-2 μM), A. flavus(MIC = 2-4 μM), A. fumigatus(MIC = 2-4 μM).
- [Ref.26028561] Drug-susceptible gram-negative bacteria: E. coli (MIC = 1 μM), P. aeruginosa (MIC = 2 μM);
- Drug-susceptible gram-positive bacteria: S. aureus (MIC = 2 μM), B. subtilis(MIC = 2 μM);
- Drug-resistant bacteria: P. aeruginosa DRPA-001 (MIC = 1 μM), P. aeruginosa DRPA-002 (MIC = 1 μM), P. aeruginosa DRPA-003 (MIC = 1 μM), P. aeruginosa DRPA-004 (MIC = 1 μM), P. aeruginosa DRPA-005 (MIC = 2 μM), S. aureus DRSA-001 (MIC = 2 μM), S. aureus DRSA-002 (MIC = 2 μM), S. aureus DRSA-003 (MIC = 1 μM), S. aureus DRSA-004 (MIC = 2 μM), S. aureus DRSA-005 (MIC = 2 μM).
-
Hemolytic Activity
-
- [Ref.26028561] HPA3NT3 and melittin revealed the hemolysis of 68.2% and 100% at 250 μM, respectively, but Hn-Mc was 1.1% at the same concentration
-
Cytotoxicity
-
- [Ref.26028561] The IC50 of HPA3NT3, melittin and Hn-Mc against HaCaT cells were 49.5, 2.8 and 357.5 μM, respectively
-
Binding Target
- Mitochondria (possibily)
Structure Information
-
Linear/Cyclic
- Linear
-
N-terminal Modification
- Free
-
C-terminal Modification
- Amidation
-
Nonterminal Modifications and Unusual Amino Acids
- None
-
Stereochemistry
- L
-
Structure
- Random coil or α-helical structure
-
Structure Description
- Hn-Mc adopted a random coil structure in the SUVs containing a mammalian membrane and in the buffer; however, it adopted an α-helical structure in SUVs containing a fungal membrane, indicating that it was not bound to the mammalian membrane but rather adhered to or inserted in the fungal membrane.
-
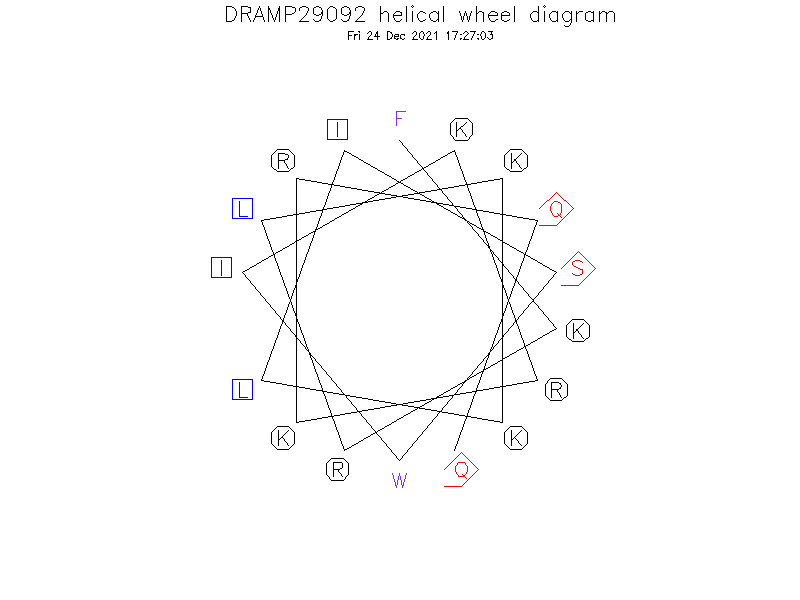
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP29092.
Physicochemical Information
-
Formula
- C105H182N34O21
Absent Amino Acids
- ACDEGHMNPTVY
Common Amino Acids
- K
Mass
- 2256.82
PI
- 12.32
Basic Residues
- 8
Acidic Residues
- 0
Hydrophobic Residues
- 6
Net Charge
- +8
-
Boman Index
- -6200
Hydrophobicity
- -1.312
Aliphatic Index
- 91.76
Half Life
-
- Mammalian:1.1 hour
- Yeast:3 min
- E.coli:2 min
Extinction Coefficient Cystines
- 5500
Absorbance 280nm
- 343.75
Polar Residues
- 1
DRAMP29092
Comments Information
Comment
- Hn-Mc has a high affinity for the fungal plasma membrane and induces apoptosis in fungal cells, and provide guidance for the development of new antifungal agents. Hn-Mc is an excellent model peptide with potent antibacterial activity and non-cytotoxicity.
Literature Information
- ·Literature 1
-
Title
- Antifungal Effect of A Chimeric Peptide Hn-Mc against Pathogenic Fungal Strains.
-
Pubmed ID
- 32731574
-
Reference
- Antibiotics (Basel). 2020 Jul 28;9(8):454. doi: 10.3390/antibiotics9080454.
-
Author
- Kim JY, Park SC, Noh G, Kim H, Yoo SH, Kim IR, Lee JR, Jang MK.
- ·Literature 2
-
Title
- Novel chimeric peptide with enhanced cell specificity and anti-inflammatory activity.
-
Pubmed ID
- 26028561
-
Reference
- Biochem Biophys Res Commun. 2015 Jul 31;463(3):322-8. doi: 10.1016/j.bbrc.2015.05.063. Epub 2015 May 29.
-
Author
- Young-Min Kim, Nam-Hong Kim, Jong-Wan Lee, Jin-Sun Jang, Yung-Hoon Park, Seong-Cheol Park, Mi-Kyeong Jang