General Information
-
DRAMP ID
- DRAMP29104
-
Peptide Name
- Mastoparan-C(MP-C)
-
Source
- Vespa crabro
-
Family
- Not found
-
Gene
- N/A
-
Sequence
- LNLKALLAVAKKIL
-
Sequence Length
- 14
-
UniProt Entry
- P01516
-
Protein Existence
- Protein level
Activity Information
-
Biological Activity
- Antimicrobial, Antibacterial,Anti-Gram+, Anti-Gram-, Anti-cancer,Antifungal
-
Target Organism
-
- [Ref.29904274]Gram-positive bacteria:Staphylococcus aureus NCTC 10788(MIC=2 µM); Staphylococcus aureus NCTC 10788(MBC=2 µM); Staphylococcus aureus ATCC 12493(MRSA,MIC=4 µM); Staphylococcus aureus ATCC 12493(MRSA,MBC=4 µM); Enterococcus faecalis NCTC 12697(MIC=8 µM); Enterococcus faecalis NCTC 12697(MBC=8 µM);
- Gram-negative bacteria:Escherichia coli NCTC 10418(MIC=4 µM); Escherichia coli NCTC 10418(MBC=8 µM); Pseudomonas aeruginosa ATCC 27853(MIC=8 µM); Pseudomonas aeruginosa ATCC 27853(MBC=16 µM); Fungi:Candida albicans NCTC 1467(MIC=4 µM); Candida albicans NCTC 1467(MBC=4 µM);
- Cancer:Human squamous lung carcinoma NCI-H157(IC50=13.57 µM);Human breast adenocarcinoma MDA-MB-435S(IC50=27.70 µM);Human prostate adenocarcinoma PC-3(IC50=6.29 µM);Human glioblastoma U251-MG(IC50=36.65 µM); Human breast adenocarcinoma MCF-7(IC50=25.27 µM).
- [Ref.33285267]Gram-positive bacteria:Staphylococcus aureus ATCC 25923(MIC=4 µM); Bacillus subtilis ATCC 23857(MIC=4 µM);
- Gram-negative bacterial:Escherichia coli ATCC 25922(4.5 mM KCl and 0.004 mM FeCl3,MIC=4 µM); Pseudomonas aeruginosa ATCC 9027(4.5 mM KCl,MIC=8 µM); Klebsiella pneumoniae ATCC 700603(MIC=8 µM); Escherichia coli ATCC 25922(NaCl/MgCl2=150mM/1mM,MIC=4 µM); Pseudomonas aeruginosa ATCC 9027(NaCl/MgCl2=150mM/1mM,MIC=16 µM); Pseudomonas aeruginosa ATCC 9027(0.004mM FeCl3,MIC=4 µM);Escherichia coli(Rifampin-resistant strain,MIC=4 µM).
-
Hemolytic Activity
-
- [Ref.29904274]50% hemolysis against horse erythrocytes at 40.11 µM.
- [Ref.33285267]10% hemolysis against mouse erythrocytes at 64 µM; 60% hemolysis against mouse erythrocytes at 256 µM.
-
Cytotoxicity
-
- [Ref.29904274]Cytotoxicity:Human microvascular endothelial cells HMEC-1(IC50=57.15 µM).
- [Ref.33285267]90% Killing against Human embryonic kidney HEK293T cells at 128 µM; Human embryonic kidney HEK293T cells(IC50=16 µM).
-
Binding Target
- liposomes
Structure Information
-
Linear/Cyclic
- Linear
-
N-terminal Modification
- Free
-
C-terminal Modification
- Amidation
-
Nonterminal Modifications and Unusual Amino Acids
- None
-
Stereochemistry
- L
-
Structure
- Alpha helix,random coil
-
Structure Description
- The CD spectra in water adopts a random-coil conformation,24.85% α-helix was determined in 50% TFE.
-
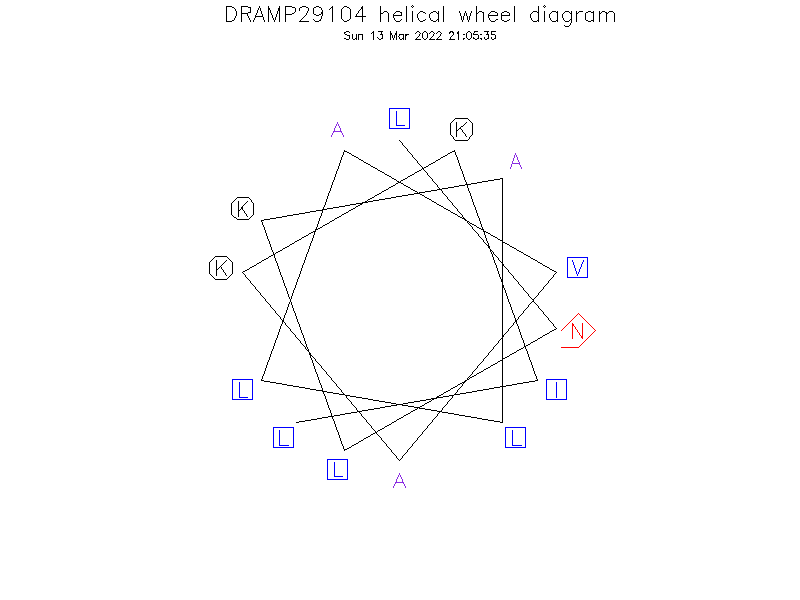
Helical Wheel Diagram
-
PDB ID
- None
-
Predicted Structure
- There is no predicted structure for DRAMP29104.
Physicochemical Information
-
Formula
- C72H134N18O16
Absent Amino Acids
- CDEFGHMPQRSTWY
Common Amino Acids
- L
Mass
- 1507.97
PI
- 10.3
Basic Residues
- 3
Acidic Residues
- 0
Hydrophobic Residues
- 10
Net Charge
- +3
-
Boman Index
- 1570
Hydrophobicity
- 1.279
Aliphatic Index
- 209.29
Half Life
-
- Mammalian:5.5 hour
- Yeast:3 min
- E.coli:2 min
Extinction Coefficient Cystines
- 0
Absorbance 280nm
- 0
Polar Residues
- 1
DRAMP29104
Comments Information
Function
- The peptide was tolerant in the presence of physiological salts.MP-C lost antimicrobial activities after incubation with 20 μg/mL trypsin or chymotrypsin,indicating moderate protease resistance.
Literature Information
- ·Literature 1
-
Title
- Newly designed antimicrobial peptides with potent bioactivity and enhanced cell selectivity prevent and reverse rifampin resistance in Gram-negative bacteria.
-
Pubmed ID
- 33285267
-
Reference
- Eur J Pharm Sci. 2021 Mar 1;158:105665.
-
Author
- Zhu N, Zhong C, Liu T, Zhu Y, Gou S, Bao H, Yao J, Ni J.
- ·Literature 2
-
Title
- Evaluation of the bioactivity of a mastoparan peptide from wasp venom and of its analogues designed through targeted engineering.
-
Pubmed ID
- 29904274
-
Reference
- Int J Biol Sci. 2018 Apr 25;14(6):599-607.
-
Author
- Chen X, Zhang L, Wu Y, Wang L, Ma C, Xi X, Bininda-Emonds ORP, Shaw C, Chen T, Zhou M.